Powerful Machine-Learning Approach Helps Identify Hundreds of New Potential COVID-19 Drugs
|
By HospiMedica International staff writers Posted on 14 Aug 2020 |
Illustration
Scientists at the University of California, Riverside (Riverside, CA, USA), have used machine learning to identify hundreds of new potential drugs that could help treat COVID-19, the disease caused by the novel coronavirus, or SARS-CoV-2.
The drug discovery pipeline is a type of computational strategy linked to artificial intelligence - a computer algorithm that learns to predict activity through trial and error, improving over time. The scientists used small numbers of previously known ligands for 65 human proteins that are known to interact with SARS-CoV-2 proteins and generated machine learning models for each of the human proteins. These models are trained to identify new small molecule inhibitors and activators, the ligands, simply from their 3-D structures. The scientists were thus able to create a database of chemicals whose structures were predicted as interactors of the 65 protein targets and also evaluate the chemicals for safety.
The scientists used their machine learning models to screen more than 10 million commercially available small molecules from a database comprised of 200 million chemicals, and identified the best-in-class hits for the 65 human proteins that interact with SARS-CoV-2 proteins. Taking it a step further, they identified compounds among the hits that are already FDA approved, such as drugs and compounds used in food. They also used the machine learning models to compute toxicity, which helped them reject potentially toxic candidates. This helped them prioritize the chemicals that were predicted to interact with SARS-CoV-2 targets. Their method allowed them to not only identify the highest scoring candidates with significant activity against a single human protein target, but also find a few chemicals that were predicted to inhibit two or more human protein targets.
“There is an urgent need to identify effective drugs that treat or prevent COVID-19. We have developed a drug discovery pipeline that identified several candidates,” said Anandasankar Ray, a professor of molecular, cell, and systems biology who led the research. “As a result, drug candidate pipelines, such as the one we developed, are extremely important to pursue as a first step toward systematic discovery of new drugs for treating COVID-19. Existing FDA-approved drugs that target one or more human proteins important for viral entry and replication are currently high priority for repurposing as new COVID-19 drugs. The demand is high for additional drugs or small molecules that can interfere with both entry and replication of SARS-CoV-2 in the body. Our drug discovery pipeline can help.”
“Our database can serve as a resource for rapidly identifying and testing novel, safe treatment strategies for COVID-19 and other diseases where the same 65 target proteins are relevant,” said Joel Kowalewski, a graduate student in Ray’s lab. “While the COVID-19 pandemic was what motivated us, we expect our predictions from more than 10 million chemicals will accelerate drug discovery in the fight against not only COVID-19 but also a number of other diseases.”
Related Links:
University of California, Riverside
The drug discovery pipeline is a type of computational strategy linked to artificial intelligence - a computer algorithm that learns to predict activity through trial and error, improving over time. The scientists used small numbers of previously known ligands for 65 human proteins that are known to interact with SARS-CoV-2 proteins and generated machine learning models for each of the human proteins. These models are trained to identify new small molecule inhibitors and activators, the ligands, simply from their 3-D structures. The scientists were thus able to create a database of chemicals whose structures were predicted as interactors of the 65 protein targets and also evaluate the chemicals for safety.
The scientists used their machine learning models to screen more than 10 million commercially available small molecules from a database comprised of 200 million chemicals, and identified the best-in-class hits for the 65 human proteins that interact with SARS-CoV-2 proteins. Taking it a step further, they identified compounds among the hits that are already FDA approved, such as drugs and compounds used in food. They also used the machine learning models to compute toxicity, which helped them reject potentially toxic candidates. This helped them prioritize the chemicals that were predicted to interact with SARS-CoV-2 targets. Their method allowed them to not only identify the highest scoring candidates with significant activity against a single human protein target, but also find a few chemicals that were predicted to inhibit two or more human protein targets.
“There is an urgent need to identify effective drugs that treat or prevent COVID-19. We have developed a drug discovery pipeline that identified several candidates,” said Anandasankar Ray, a professor of molecular, cell, and systems biology who led the research. “As a result, drug candidate pipelines, such as the one we developed, are extremely important to pursue as a first step toward systematic discovery of new drugs for treating COVID-19. Existing FDA-approved drugs that target one or more human proteins important for viral entry and replication are currently high priority for repurposing as new COVID-19 drugs. The demand is high for additional drugs or small molecules that can interfere with both entry and replication of SARS-CoV-2 in the body. Our drug discovery pipeline can help.”
“Our database can serve as a resource for rapidly identifying and testing novel, safe treatment strategies for COVID-19 and other diseases where the same 65 target proteins are relevant,” said Joel Kowalewski, a graduate student in Ray’s lab. “While the COVID-19 pandemic was what motivated us, we expect our predictions from more than 10 million chemicals will accelerate drug discovery in the fight against not only COVID-19 but also a number of other diseases.”
Related Links:
University of California, Riverside
Latest AI News
- Machine Learning Approach Enhances Liver Cancer Risk Stratification
- New AI Approach Monitors Brain Health Using Passive Wearable Data
- AI Tool Maps Early Risk Patterns in Bloodstream Infections
- AI Model Identifies Rare Endocrine Disorder from Hand Images
- AI Tool Promises to Reduce Length of Hospital Stays and Free Up Beds
- Machine Learning Model Cuts Canceled Liver Transplants By 60%
Channels
Artificial Intelligence
view channel
Machine Learning Approach Enhances Liver Cancer Risk Stratification
Hepatocellular carcinoma, the most common form of primary liver cancer, is often detected late despite targeted surveillance programs. Current screening guidelines emphasize patients with known cirrhosis,... Read more
New AI Approach Monitors Brain Health Using Passive Wearable Data
Brain health spans cognitive and emotional functions and can fluctuate even in adults without diagnosed disease. Detecting early changes remains difficult in routine care and burdens specialty services... Read moreCritical Care
view channel
Automated IV Labeling Solution Improves Infusion Safety and Efficiency
Medication administration in high-acuity settings is often complicated by multiple concurrent infusions, making accurate line identification essential. In a 10-hospital intensive care unit study, 60% of... Read more
First-Of-Its-Kind AI Tool Detects Pulmonary Hypertension from Standard ECGs
Pulmonary hypertension is a progressive, life‑threatening disease that is frequently missed early because symptoms such as dyspnea are nonspecific and diagnostic delays can exceed two years.... Read moreSurgical Techniques
view channel
Continuous Monitoring with Wearables Enhances Postoperative Patient Safety
Postoperative hypoxemia on general surgical wards is common and often missed by intermittent vital sign checks. Undetected low oxygen levels can delay recovery and raise the risk of complications that... Read more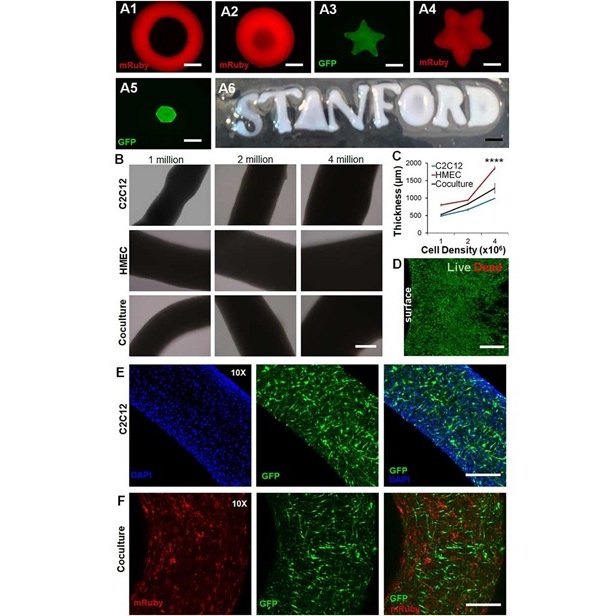
New Approach Enables Customized Muscle Tissue Without Biomaterial Scaffolds
Volumetric muscle loss is a traumatic loss of skeletal muscle that often leads to permanent functional impairment and limited reconstructive options. Current experimental strategies struggle to deliver... Read morePatient Care
view channel
Wearable Sleep Data Predict Adherence to Pulmonary Rehabilitation
Chronic obstructive pulmonary disease (COPD) is a long-term lung disorder that makes breathing difficult and often disturbs sleep, reducing energy for daily activities. Limited engagement in pulmonary... Read more
Revolutionary Automatic IV-Line Flushing Device to Enhance Infusion Care
More than 80% of in-hospital patients receive intravenous (IV) therapy. Every dose of IV medicine delivered in a small volume (<250 mL) infusion bag should be followed by subsequent flushing to ensure... Read moreHealth IT
view channel
EMR-Based Tool Predicts Graft Failure After Kidney Transplant
Kidney transplantation offers patients with end-stage kidney disease longer survival and better quality of life than dialysis, yet graft failure remains a major challenge. Although a successful transplant... Read more
Printable Molecule-Selective Nanoparticles Enable Mass Production of Wearable Biosensors
The future of medicine is likely to focus on the personalization of healthcare—understanding exactly what an individual requires and delivering the appropriate combination of nutrients, metabolites, and... Read moreBusiness
view channel