Antibiotic Management Strategies Fail to Curb Resistance
|
By HospiMedica International staff writers Posted on 31 Jan 2017 |
A new study concludes that neither of the two current antibiotic regimens employed in hospitals can effectively mitigate antibiotic resistance.
Researchers at the University of Exeter, the University of Sydney, and other institutions conducted a theoretical mathematical study to try and explain why recent clinical trials are unable to resolve the highly controversial issue of which strategy to combat antibiotic resistance is superior - antibiotic cycling or mixing. Cycling implements the restriction and prioritization of different antibiotics at different times in hospitals; in antibiotic mixing, appropriate antibiotics are allocated to patients in a random fashion.
The researchers concluded, based on almost four million patient days of treatment tracked in the clinical trials, that the literature that showed support for the theoretical optimality of mixing was misplaced, and that clinical datasets cannot choose between the two. Their analysis is consistent with an emerging pattern that shows that neither cycling nor mixing are better than the other at mitigating pathogen selection for antibiotic resistance. The study was published on January 17, 2017, in Molecular Biology and Evolution.
“Mathematically speaking, it was very clear early in this study that antibiotic mixing was not the optimal way of allocating antibiotic to patients, yet this is what some clinicians have come believe,” said lead author Professor Robert Beardmore, PhD, of Exeter University. “But communicating this was difficult, given the complexity of the mathematical ideas. In the end, the real mathematical optimum is little more than common sense: get the right drugs to the right patients as soon as possible.”
“It is clear that information-rich, personalized protocols can outperform antibiotic cycling and mixing in mathematical models, but even this conclusion can depend on nuanced model circumstances,” said study co-author Rafael Pena-Miller, PhD, of Exeter University. “In the doomsday scenario that multi-drug resistance is endemic and present in every infection before the patient begins treatment, it matters little which treatment patients are given. But before that stark situation arises, targeting appropriate treatment at as many individuals as possible outperforms both mixing and cycling.”
“Personalized medicine is rapidly becoming a reality with dramatic increases in the availability of clinical testing at the point of care,” added senior author Professor Jon Iredell, MD, of the University of Sydney. “Antibiotic use in severe infection remains one of the most powerful interventions in medicine, and intelligent use of antibiotics is essential to optimize immediate patient outcomes and to preserve long-term benefits.”
The researchers therefore recommend that individualized treatments, both pathogen-specific and patient-specific, may be a necessity to properly optimize antibiotic use. By using computer models to study different personalized medicine scenarios, they advocate the use of devices that target infections based on rapid diagnoses of the pathogen responsible for the infection from molecular signatures or blood cultures.
Researchers at the University of Exeter, the University of Sydney, and other institutions conducted a theoretical mathematical study to try and explain why recent clinical trials are unable to resolve the highly controversial issue of which strategy to combat antibiotic resistance is superior - antibiotic cycling or mixing. Cycling implements the restriction and prioritization of different antibiotics at different times in hospitals; in antibiotic mixing, appropriate antibiotics are allocated to patients in a random fashion.
The researchers concluded, based on almost four million patient days of treatment tracked in the clinical trials, that the literature that showed support for the theoretical optimality of mixing was misplaced, and that clinical datasets cannot choose between the two. Their analysis is consistent with an emerging pattern that shows that neither cycling nor mixing are better than the other at mitigating pathogen selection for antibiotic resistance. The study was published on January 17, 2017, in Molecular Biology and Evolution.
“Mathematically speaking, it was very clear early in this study that antibiotic mixing was not the optimal way of allocating antibiotic to patients, yet this is what some clinicians have come believe,” said lead author Professor Robert Beardmore, PhD, of Exeter University. “But communicating this was difficult, given the complexity of the mathematical ideas. In the end, the real mathematical optimum is little more than common sense: get the right drugs to the right patients as soon as possible.”
“It is clear that information-rich, personalized protocols can outperform antibiotic cycling and mixing in mathematical models, but even this conclusion can depend on nuanced model circumstances,” said study co-author Rafael Pena-Miller, PhD, of Exeter University. “In the doomsday scenario that multi-drug resistance is endemic and present in every infection before the patient begins treatment, it matters little which treatment patients are given. But before that stark situation arises, targeting appropriate treatment at as many individuals as possible outperforms both mixing and cycling.”
“Personalized medicine is rapidly becoming a reality with dramatic increases in the availability of clinical testing at the point of care,” added senior author Professor Jon Iredell, MD, of the University of Sydney. “Antibiotic use in severe infection remains one of the most powerful interventions in medicine, and intelligent use of antibiotics is essential to optimize immediate patient outcomes and to preserve long-term benefits.”
The researchers therefore recommend that individualized treatments, both pathogen-specific and patient-specific, may be a necessity to properly optimize antibiotic use. By using computer models to study different personalized medicine scenarios, they advocate the use of devices that target infections based on rapid diagnoses of the pathogen responsible for the infection from molecular signatures or blood cultures.
Latest Critical Care News
- Stretchable Microneedles to Help In Accurate Tracking of Abnormalities and Identifying Rapid Treatment
- Machine Learning Tool Identifies Rare, Undiagnosed Immune Disorders from Patient EHRs
- On-Skin Wearable Bioelectronic Device Paves Way for Intelligent Implants
- First-Of-Its-Kind Dissolvable Stent to Improve Outcomes for Patients with Severe PAD
- AI Brain-Age Estimation Technology Uses EEG Scans to Screen for Degenerative Diseases
- Wheeze-Counting Wearable Device Monitors Patient's Breathing In Real Time
- Wearable Multiplex Biosensors Could Revolutionize COPD Management
- New Low-Energy Defibrillation Method Controls Cardiac Arrhythmias
- New Machine Learning Models Help Predict Heart Disease Risk in Women
- Deep-Learning Model Predicts Arrhythmia 30 Minutes before Onset
- Breakthrough Technology Combines Detection and Treatment of Nerve-Related Disorders in Single Procedure
- Plasma Irradiation Promotes Faster Bone Healing
- New Device Treats Acute Kidney Injury from Sepsis
- Study Confirms Safety of DCB-Only Strategy for Treating De Novo Left Main Coronary Artery Disease
- Revascularization Improves Quality of Life for Patients with Chronic Limb Threatening Ischemia
- AI-Driven Prediction Models Accurately Predict Critical Care Patient Deterioration
Channels
Artificial Intelligence
view channel
AI-Powered Algorithm to Revolutionize Detection of Atrial Fibrillation
Atrial fibrillation (AFib), a condition characterized by an irregular and often rapid heart rate, is linked to increased risks of stroke and heart failure. This is because the irregular heartbeat in AFib... Read more
AI Diagnostic Tool Accurately Detects Valvular Disorders Often Missed by Doctors
Doctors generally use stethoscopes to listen for the characteristic lub-dub sounds made by heart valves opening and closing. They also listen for less prominent sounds that indicate problems with these valves.... Read moreSurgical Techniques
view channel
World’s Smallest Laser Probe for Brain Procedures Facilitates Ablation of Full Range of Targets
A new probe enhances the ablation capabilities for a broad spectrum of oncology and epilepsy targets, including pediatric applications, by incorporating advanced laser and cooling technologies to support... Read more.jpg)
Artificial Intelligence Broadens Diagnostic Abilities of Conventional Coronary Angiography
Coronary angiography is an essential diagnostic tool used globally to identify coronary artery disease (CAD), with millions of procedures conducted annually. Traditionally, data from coronary angiograms... Read moreAI-Powered Surgical Visualization Tool Supports Surgeons' Visual Recognition in Real Time
Connective tissue serves as an essential landmark in surgical navigation, often referred to as the "dissection plane" or "holy plane." Its accurate identification is vital for achieving safe and effective... Read moreHandheld Device for Fluorescence-Guided Surgery a Game Changer for Removal of High-Grade Glioma Brain Tumors
Grade III or IV gliomas are among the most common and deadly brain tumors, with around 20,000 cases annually in the U.S. and 1.2 million globally. These tumors are very aggressive and tend to infiltrate... Read morePatient Care
view channelFirst-Of-Its-Kind Portable Germicidal Light Technology Disinfects High-Touch Clinical Surfaces in Seconds
Reducing healthcare-acquired infections (HAIs) remains a pressing issue within global healthcare systems. In the United States alone, 1.7 million patients contract HAIs annually, leading to approximately... Read more
Surgical Capacity Optimization Solution Helps Hospitals Boost OR Utilization
An innovative solution has the capability to transform surgical capacity utilization by targeting the root cause of surgical block time inefficiencies. Fujitsu Limited’s (Tokyo, Japan) Surgical Capacity... Read more
Game-Changing Innovation in Surgical Instrument Sterilization Significantly Improves OR Throughput
A groundbreaking innovation enables hospitals to significantly improve instrument processing time and throughput in operating rooms (ORs) and sterile processing departments. Turbett Surgical, Inc.... Read moreHealth IT
view channel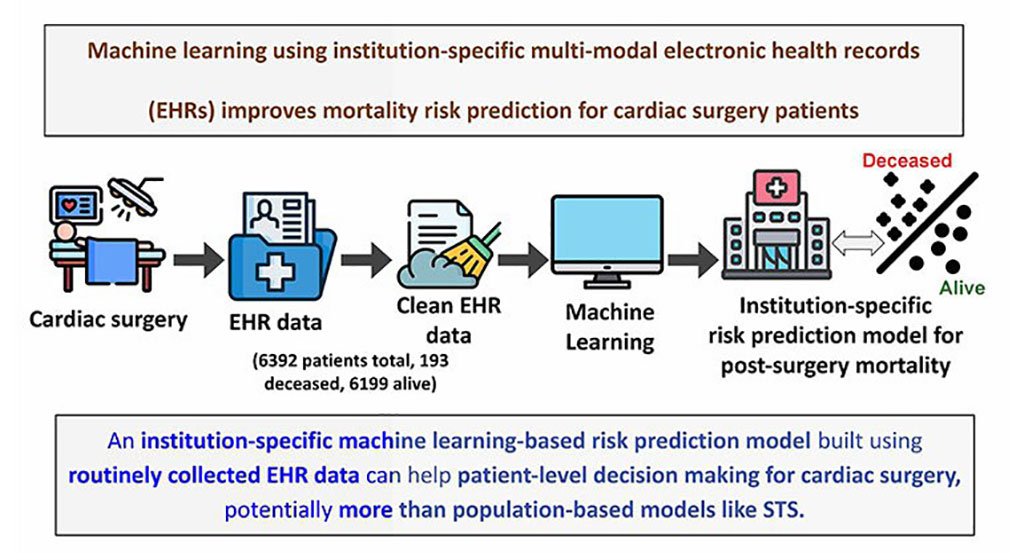
Machine Learning Model Improves Mortality Risk Prediction for Cardiac Surgery Patients
Machine learning algorithms have been deployed to create predictive models in various medical fields, with some demonstrating improved outcomes compared to their standard-of-care counterparts.... Read more
Strategic Collaboration to Develop and Integrate Generative AI into Healthcare
Top industry experts have underscored the immediate requirement for healthcare systems and hospitals to respond to severe cost and margin pressures. Close to half of U.S. hospitals ended 2022 in the red... Read more
AI-Enabled Operating Rooms Solution Helps Hospitals Maximize Utilization and Unlock Capacity
For healthcare organizations, optimizing operating room (OR) utilization during prime time hours is a complex challenge. Surgeons and clinics face difficulties in finding available slots for booking cases,... Read more
AI Predicts Pancreatic Cancer Three Years before Diagnosis from Patients’ Medical Records
Screening for common cancers like breast, cervix, and prostate cancer relies on relatively simple and highly effective techniques, such as mammograms, Pap smears, and blood tests. These methods have revolutionized... Read morePoint of Care
view channel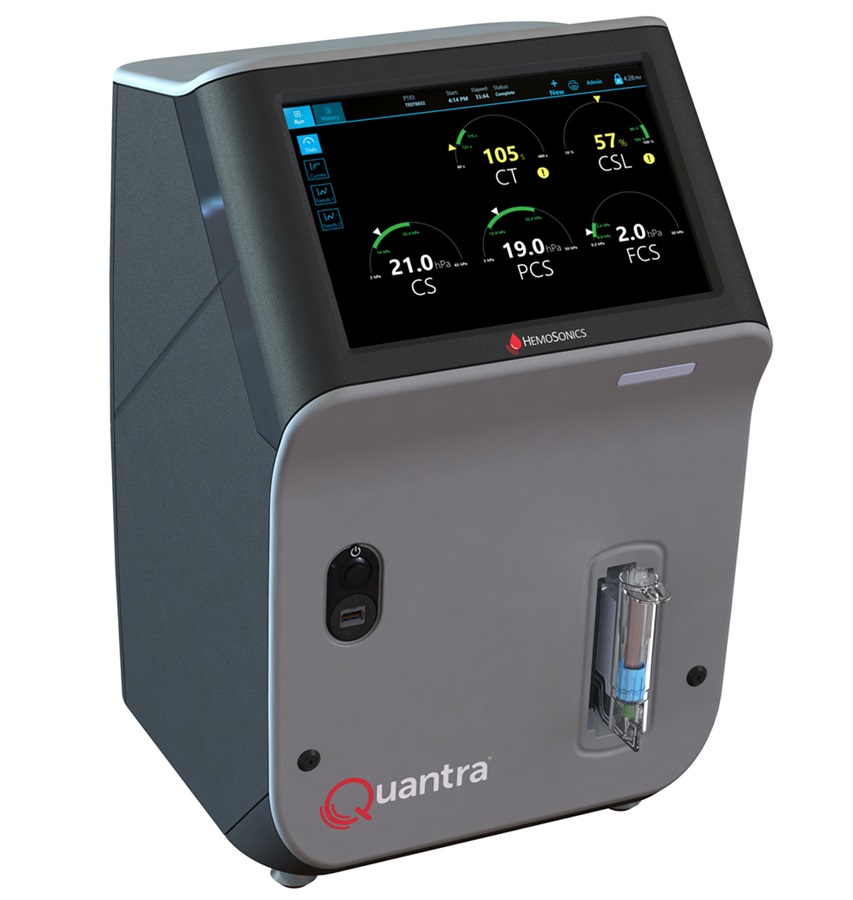
Critical Bleeding Management System to Help Hospitals Further Standardize Viscoelastic Testing
Surgical procedures are often accompanied by significant blood loss and the subsequent high likelihood of the need for allogeneic blood transfusions. These transfusions, while critical, are linked to various... Read more
Point of Care HIV Test Enables Early Infection Diagnosis for Infants
Early diagnosis and initiation of treatment are crucial for the survival of infants infected with HIV (human immunodeficiency virus). Without treatment, approximately 50% of infants who acquire HIV during... Read more
Whole Blood Rapid Test Aids Assessment of Concussion at Patient's Bedside
In the United States annually, approximately five million individuals seek emergency department care for traumatic brain injuries (TBIs), yet over half of those suspecting a concussion may never get it checked.... Read more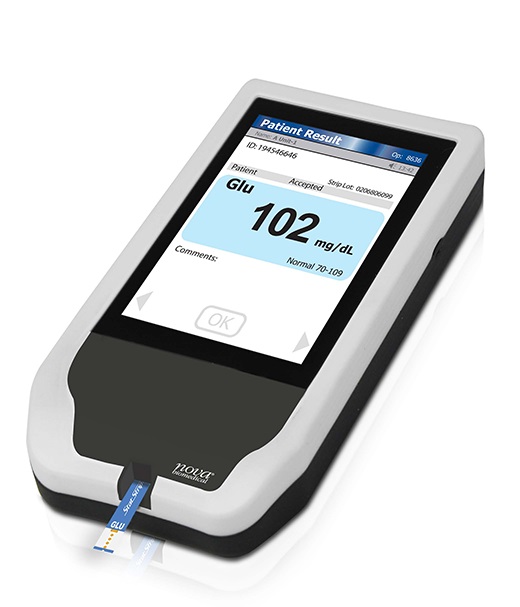
New Generation Glucose Hospital Meter System Ensures Accurate, Interference-Free and Safe Use
A new generation glucose hospital meter system now comes with several features that make hospital glucose testing easier and more secure while continuing to offer accuracy, freedom from interference, and... Read moreBusiness
view channel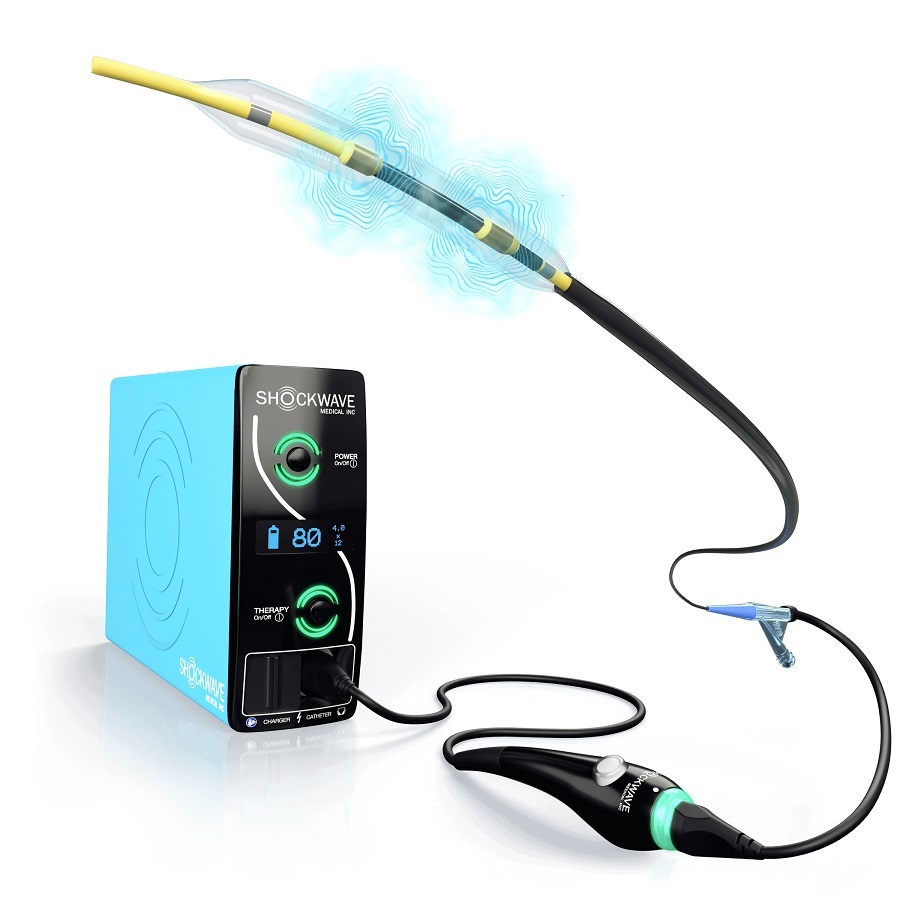
Johnson & Johnson Acquires Cardiovascular Medical Device Company Shockwave Medical
Johnson & Johnson (New Brunswick, N.J., USA) and Shockwave Medical (Santa Clara, CA, USA) have entered into a definitive agreement under which Johnson & Johnson will acquire all of Shockwave’s... Read more