Deep-Learning Model that Can Predict How Human Genes and Medicines Will Interact Identifies Promising COVID-19 Treatments
|
By HospiMedica International staff writers Posted on 03 Feb 2021 |

Image: The computer model can predict how human genes and medicines will interact (Photo courtesy of Getty Images)
A new deep-learning model that can predict how human genes and medicines will interact has identified at least 10 compounds that may hold promise as treatments for COVID-19.
The computer model, which the researchers call “DeepCE” - pronounced “Deep Sea” was created by computer scientists at The Ohio State University (Columbus, OH, USA) and helps find drug repurposing candidates for new diseases. All but two of the drugs identified as potential COVID-19 treatments by the deep-learning model are still considered investigational and are being tested for effectiveness against hepatitis C, fungal disease, cancer and heart disease. The list also includes the approved drugs cyclosporine, an immunosuppressant that prevents transplant organ rejection, and anidulafungin, an antifungal agent.
Much more work needs to be done before any of these medications would be confirmed as safe and effective treatments for people infected with SARS-CoV-2. But by using artificial intelligence to arrive at these options, the scientists have saved pharmaceutical and clinical researchers the time and money it would take to search for potential COVID-19 drugs on a piecemeal basis. The researchers have noted that a few of the repurposing candidates the model generated have already been studied for their potential use in COVID-19 patients.
To make predictions about how genes and medicines will interact and yield drug repurposing candidates, DeepCE relies on two primary sources of publicly available data: L1000, a National Institutes of Health-funded repository of human cell-line data showing how gene expression changes in response to drugs, and DrugBank, which contains information on the chemical structures and other details on about 11,000 approved and investigational drugs.
L1000 displays side-by-side cell-line comparisons of standard gene expression activity with gene expression changes produced by interactions with specific drugs. The cell lines represent diseases, such as melanoma, and organs, like kidneys and lungs. It is an ongoing project, with data being added as experiments in animals or humans supplement the gene expression profiles produced in cell-line experiments.
The Ohio State researchers trained the DeepCE model by running all of the L1000 data through an algorithm against specific chemical compounds and their dosages. To fill in data gaps, the model converts chemical compound descriptions into figures, allowing for automatic consideration of their separate components’ effects on genes. And for genes not represented in L1000, the team used a deep learning approach called an “attention mechanism” to increase the model’s “learned” sample of gene-chemical compound interactions, which improves the framework’s performance.
The team applied DeepCE’s gene expression prediction matrix - focusing on data from lung and airway cell lines and the entire DrugBank catalog of compounds - to the genetic information provided from the early COVID-19 papers and additional government data. The COVID-19 data demonstrated how human gene expression had responded to being infected with SARS-CoV-2, creating a “disease signature.”
“When no one has any information on a new disease, this model shows how artificial intelligence can help solve the problem of how to consider a potential treatment,” said senior author Ping Zhang, assistant professor of computer science and engineering and biomedical informatics at The Ohio State University. “Great minds think alike - some lead compounds identified by machine intelligence coincide with later discoveries by human intelligence.”
Related Links:
The Ohio State University
The computer model, which the researchers call “DeepCE” - pronounced “Deep Sea” was created by computer scientists at The Ohio State University (Columbus, OH, USA) and helps find drug repurposing candidates for new diseases. All but two of the drugs identified as potential COVID-19 treatments by the deep-learning model are still considered investigational and are being tested for effectiveness against hepatitis C, fungal disease, cancer and heart disease. The list also includes the approved drugs cyclosporine, an immunosuppressant that prevents transplant organ rejection, and anidulafungin, an antifungal agent.
Much more work needs to be done before any of these medications would be confirmed as safe and effective treatments for people infected with SARS-CoV-2. But by using artificial intelligence to arrive at these options, the scientists have saved pharmaceutical and clinical researchers the time and money it would take to search for potential COVID-19 drugs on a piecemeal basis. The researchers have noted that a few of the repurposing candidates the model generated have already been studied for their potential use in COVID-19 patients.
To make predictions about how genes and medicines will interact and yield drug repurposing candidates, DeepCE relies on two primary sources of publicly available data: L1000, a National Institutes of Health-funded repository of human cell-line data showing how gene expression changes in response to drugs, and DrugBank, which contains information on the chemical structures and other details on about 11,000 approved and investigational drugs.
L1000 displays side-by-side cell-line comparisons of standard gene expression activity with gene expression changes produced by interactions with specific drugs. The cell lines represent diseases, such as melanoma, and organs, like kidneys and lungs. It is an ongoing project, with data being added as experiments in animals or humans supplement the gene expression profiles produced in cell-line experiments.
The Ohio State researchers trained the DeepCE model by running all of the L1000 data through an algorithm against specific chemical compounds and their dosages. To fill in data gaps, the model converts chemical compound descriptions into figures, allowing for automatic consideration of their separate components’ effects on genes. And for genes not represented in L1000, the team used a deep learning approach called an “attention mechanism” to increase the model’s “learned” sample of gene-chemical compound interactions, which improves the framework’s performance.
The team applied DeepCE’s gene expression prediction matrix - focusing on data from lung and airway cell lines and the entire DrugBank catalog of compounds - to the genetic information provided from the early COVID-19 papers and additional government data. The COVID-19 data demonstrated how human gene expression had responded to being infected with SARS-CoV-2, creating a “disease signature.”
“When no one has any information on a new disease, this model shows how artificial intelligence can help solve the problem of how to consider a potential treatment,” said senior author Ping Zhang, assistant professor of computer science and engineering and biomedical informatics at The Ohio State University. “Great minds think alike - some lead compounds identified by machine intelligence coincide with later discoveries by human intelligence.”
Related Links:
The Ohio State University
Latest COVID-19 News
- Low-Cost System Detects SARS-CoV-2 Virus in Hospital Air Using High-Tech Bubbles
- World's First Inhalable COVID-19 Vaccine Approved in China
- COVID-19 Vaccine Patch Fights SARS-CoV-2 Variants Better than Needles
- Blood Viscosity Testing Can Predict Risk of Death in Hospitalized COVID-19 Patients
- ‘Covid Computer’ Uses AI to Detect COVID-19 from Chest CT Scans
- MRI Lung-Imaging Technique Shows Cause of Long-COVID Symptoms
- Chest CT Scans of COVID-19 Patients Could Help Distinguish Between SARS-CoV-2 Variants
- Specialized MRI Detects Lung Abnormalities in Non-Hospitalized Long COVID Patients
- AI Algorithm Identifies Hospitalized Patients at Highest Risk of Dying From COVID-19
- Sweat Sensor Detects Key Biomarkers That Provide Early Warning of COVID-19 and Flu
- Study Assesses Impact of COVID-19 on Ventilation/Perfusion Scintigraphy
- CT Imaging Study Finds Vaccination Reduces Risk of COVID-19 Associated Pulmonary Embolism
- Third Day in Hospital a ‘Tipping Point’ in Severity of COVID-19 Pneumonia
- Longer Interval Between COVID-19 Vaccines Generates Up to Nine Times as Many Antibodies
- AI Model for Monitoring COVID-19 Predicts Mortality Within First 30 Days of Admission
- AI Predicts COVID Prognosis at Near-Expert Level Based Off CT Scans
Channels
Artificial Intelligence
view channelAI Analysis of Pericardial Fat Refines Long-Term Heart Disease Risk
Accurately identifying long-term cardiovascular disease risk in asymptomatic adults remains challenging for clinicians. Missed or underestimated risk delays preventive therapy and increases the chance... Read more
Machine Learning Approach Enhances Liver Cancer Risk Stratification
Hepatocellular carcinoma, the most common form of primary liver cancer, is often detected late despite targeted surveillance programs. Current screening guidelines emphasize patients with known cirrhosis,... Read moreCritical Care
view channel
Eye Imaging AI Identifies Elevated Cardiovascular Risk
Many adults at risk for atherosclerotic cardiovascular disease are not identified until they undergo formal primary care assessment. Delayed risk recognition can postpone initiation of statins and lifestyle... Read more
Noninvasive Monitoring Device Enables Earlier Intervention in Heart Failure
Hospitalizations for heart failure with preserved ejection fraction (HFpEF) remain common because lung congestion often worsens before symptoms prompt treatment changes. Missed early decompensation... Read moreSurgical Techniques
view channel
Fiber-Form Bone Graft Expands Intraoperative Options for Spinal Fusion
Spinal and orthopedic fusion procedures often require bone graft materials that handle predictably and support bone formation. Surgeons face added complexity in difficult anatomy and challenging fusion environments.... Read more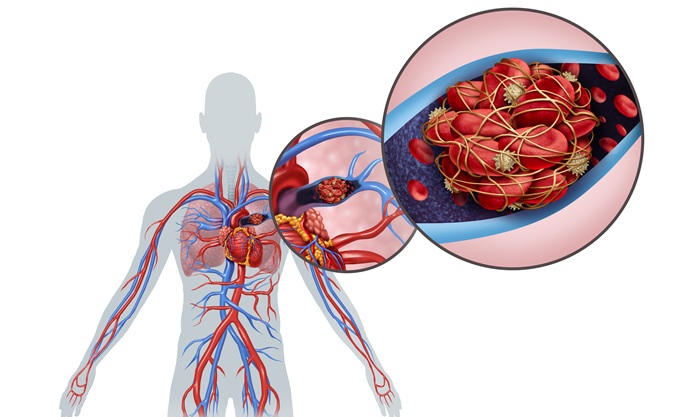
Ultrasound‑Aided Catheter Treatment Cuts Early Collapse in Pulmonary Embolism
Acute pulmonary embolism can cause rapid hemodynamic deterioration and early death in hospitalized and emergency patients. Systemic thrombolysis can dissolve clots but is limited by a high risk of major... Read morePatient Care
view channel
Wearable Sleep Data Predict Adherence to Pulmonary Rehabilitation
Chronic obstructive pulmonary disease (COPD) is a long-term lung disorder that makes breathing difficult and often disturbs sleep, reducing energy for daily activities. Limited engagement in pulmonary... Read more
Revolutionary Automatic IV-Line Flushing Device to Enhance Infusion Care
More than 80% of in-hospital patients receive intravenous (IV) therapy. Every dose of IV medicine delivered in a small volume (<250 mL) infusion bag should be followed by subsequent flushing to ensure... Read moreHealth IT
view channel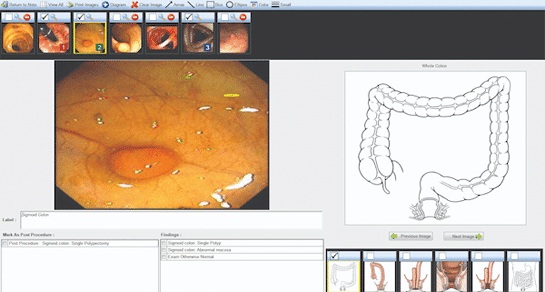
Voice-Driven AI System Enables Structured GI Procedure Documentation
Documentation during gastrointestinal (GI) procedures often competes with real-time clinical decision-making and imposes a significant cognitive burden on physicians. Manual data entry and post-procedure... Read more
EMR-Based Tool Predicts Graft Failure After Kidney Transplant
Kidney transplantation offers patients with end-stage kidney disease longer survival and better quality of life than dialysis, yet graft failure remains a major challenge. Although a successful transplant... Read more
Printable Molecule-Selective Nanoparticles Enable Mass Production of Wearable Biosensors
The future of medicine is likely to focus on the personalization of healthcare—understanding exactly what an individual requires and delivering the appropriate combination of nutrients, metabolites, and... Read moreBusiness
view channel