Wikipedia Page Views Could Predict Disease Outbreaks
|
By HospiMedica International staff writers Posted on 24 Nov 2014 |
A new study suggests that Wikipedia access data could be an effective tool for forecasting disease outbreaks up to a month in advance.
Researchers at the Los Alamos National Laboratory (NM, USA) reviewed access logs to disease-related Wikipedia pages between 2010 and 2013. They mapped the languages the information was written in, using this as an approximate measure for people's locations. Using linear statistical techniques models, the researchers then tested 14 location-disease combinations to demonstrate the feasibility of the techniques built upon the data stream, and compared the results with disease outbreak information provided by national health surveillance teams.
The researchers found three broad classes of results. In eight cases, there was a usefully close match between the model's estimate and the official data. This statistical technique allowed them to predict emerging influenza outbreaks in the United States, Poland, Japan, and Thailand, dengue fever spikes in Brazil and Thailand, and a rise in tuberculosis cases in Thailand.
In three cases, the model failed, apparently because patterns in the official data were too subtle to capture, and in a further three, the model failed apparently because the signal-to-noise ratio (SNR) in the Wikipedia data was too subtle to capture. The researchers suggested that disease incidence may also be changing too slowly to be evident in the chosen analysis period. The results also suggest that these models can be used even in places with no official data upon which to build models. The study was published on November 13, 2014, in PLoS Computational Biology.
“A global disease-forecasting system will change the way we respond to epidemics. In the same way we check the weather each morning, individuals and public health officials can monitor disease incidence and plan for the future based on today's forecast,” said lead author Sara Del Valle. “The goal of this research is to build an operational disease monitoring and forecasting system with open data and open source code. This paper shows we can achieve that goal.”
The researchers added that it is important to recognize demographic biases inherent in Wikipedia and other social internet data sources such as age, gender, and education. Most importantly, the data strongly over-represent people and places with good internet access and technology skills.
Related Links:
Los Alamos National Laboratory
Researchers at the Los Alamos National Laboratory (NM, USA) reviewed access logs to disease-related Wikipedia pages between 2010 and 2013. They mapped the languages the information was written in, using this as an approximate measure for people's locations. Using linear statistical techniques models, the researchers then tested 14 location-disease combinations to demonstrate the feasibility of the techniques built upon the data stream, and compared the results with disease outbreak information provided by national health surveillance teams.
The researchers found three broad classes of results. In eight cases, there was a usefully close match between the model's estimate and the official data. This statistical technique allowed them to predict emerging influenza outbreaks in the United States, Poland, Japan, and Thailand, dengue fever spikes in Brazil and Thailand, and a rise in tuberculosis cases in Thailand.
In three cases, the model failed, apparently because patterns in the official data were too subtle to capture, and in a further three, the model failed apparently because the signal-to-noise ratio (SNR) in the Wikipedia data was too subtle to capture. The researchers suggested that disease incidence may also be changing too slowly to be evident in the chosen analysis period. The results also suggest that these models can be used even in places with no official data upon which to build models. The study was published on November 13, 2014, in PLoS Computational Biology.
“A global disease-forecasting system will change the way we respond to epidemics. In the same way we check the weather each morning, individuals and public health officials can monitor disease incidence and plan for the future based on today's forecast,” said lead author Sara Del Valle. “The goal of this research is to build an operational disease monitoring and forecasting system with open data and open source code. This paper shows we can achieve that goal.”
The researchers added that it is important to recognize demographic biases inherent in Wikipedia and other social internet data sources such as age, gender, and education. Most importantly, the data strongly over-represent people and places with good internet access and technology skills.
Related Links:
Los Alamos National Laboratory
Latest Health IT News
- Machine Learning Model Improves Mortality Risk Prediction for Cardiac Surgery Patients
- Strategic Collaboration to Develop and Integrate Generative AI into Healthcare
- AI-Enabled Operating Rooms Solution Helps Hospitals Maximize Utilization and Unlock Capacity
- AI Predicts Pancreatic Cancer Three Years before Diagnosis from Patients’ Medical Records
- First Fully Autonomous Generative AI Personalized Medical Authorizations System Reduces Care Delay
- Electronic Health Records May Be Key to Improving Patient Care, Study Finds
- AI Trained for Specific Vocal Biomarkers Could Accurately Predict Coronary Artery Disease
- First-Ever AI Test for Early Diagnosis of Alzheimer’s to Be Expanded to Diagnosis of Parkinson’s Disease
- New Self-Learning AI-Based Algorithm Reads Electrocardiograms to Spot Unseen Signs of Heart Failure
- Autonomous Robot Performs COVID-19 Nasal Swab Tests

- Statistical Tool Predicts COVID-19 Peaks Worldwide
- Wireless-Controlled Soft Neural Implant Stimulates Brain Cells
- Tiny Polymer Stent Could Treat Pediatric Urethral Strictures
- Human Torso Simulator Helps Design Brace Innovations
- 3D Bioprinting Rebuilds the Human Heart
Channels
Artificial Intelligence
view channel
AI-Powered Algorithm to Revolutionize Detection of Atrial Fibrillation
Atrial fibrillation (AFib), a condition characterized by an irregular and often rapid heart rate, is linked to increased risks of stroke and heart failure. This is because the irregular heartbeat in AFib... Read more
AI Diagnostic Tool Accurately Detects Valvular Disorders Often Missed by Doctors
Doctors generally use stethoscopes to listen for the characteristic lub-dub sounds made by heart valves opening and closing. They also listen for less prominent sounds that indicate problems with these valves.... Read moreCritical Care
view channel
AI to Improved Diagnosis of Atrial Fibrillation
Abnormal heart rhythms frequently arise from—and contribute to—structural abnormalities in the heart. Atrial fibrillation is a specific type of abnormal rhythm that may not be consistently present, often... Read more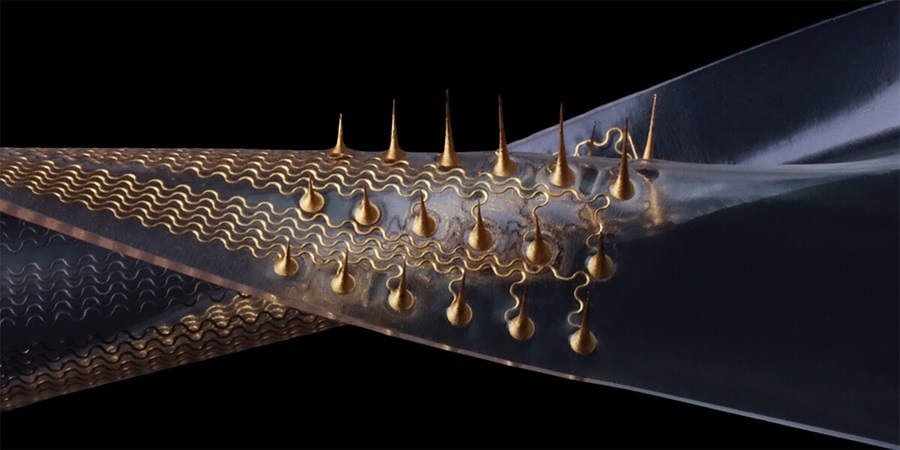
Stretchable Microneedles to Help In Accurate Tracking of Abnormalities and Identifying Rapid Treatment
The field of personalized medicine is transforming rapidly, with advancements like wearable devices and home testing kits making it increasingly easy to monitor a wide range of health metrics, from heart... Read more
Machine Learning Tool Identifies Rare, Undiagnosed Immune Disorders from Patient EHRs
Patients suffering from rare diseases often endure extensive delays in receiving accurate diagnoses and treatments, which can lead to unnecessary tests, worsening health, psychological strain, and significant... Read moreSurgical Techniques
view channel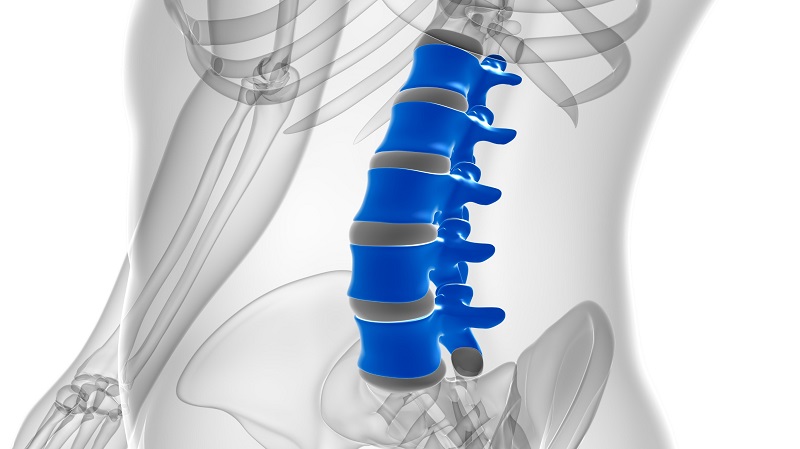
Tiny Wraparound Electronic Implants to Revolutionize Treatment of Spinal Cord Injuries
The spinal cord functions as a vital conduit, transmitting nerve impulses to and from the brain, much like a highway. When the spinal cord is damaged, this flow of information is disrupted, leading to... Read more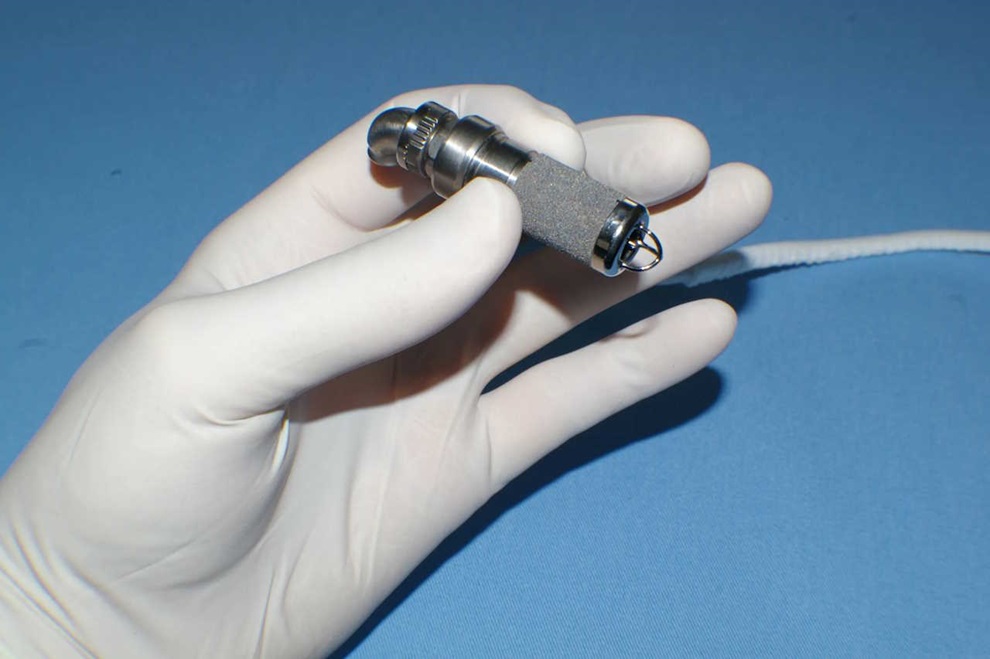
Small, Implantable Cardiac Pump to Help Children Awaiting Heart Transplant
Implantable ventricular assist devices, available for adults for over 40 years, fit inside the chest and are generally safer and easier to use than external devices. These devices enable patients to live... Read moreGastrointestinal Imaging Capsule a Game-Changer in Esophagus Surveillance and Treatment
A newly-developed gastrointestinal imaging capsule is poised to be a game-changer in esophagus surveillance and interventions. The Multifunctional Ablative Gastrointestinal Imaging Capsule (MAGIC) developed... Read more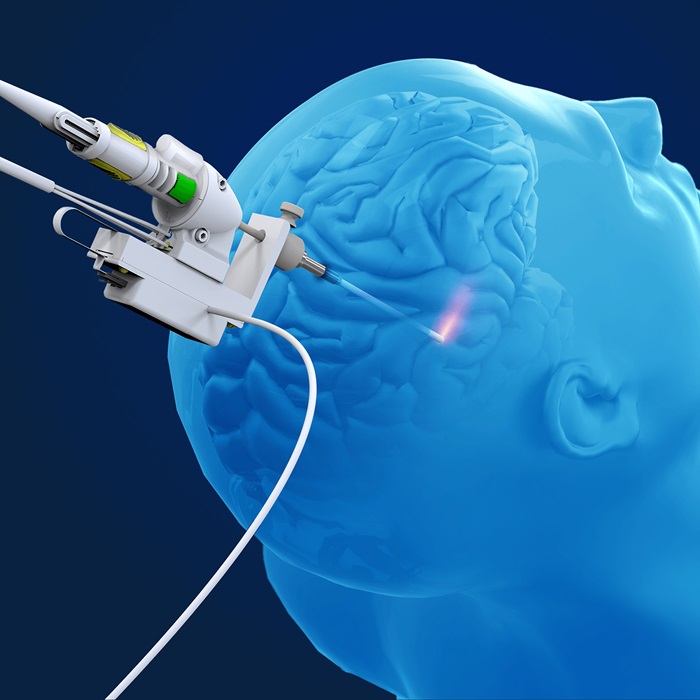
World’s Smallest Laser Probe for Brain Procedures Facilitates Ablation of Full Range of Targets
A new probe enhances the ablation capabilities for a broad spectrum of oncology and epilepsy targets, including pediatric applications, by incorporating advanced laser and cooling technologies to support... Read morePatient Care
view channelFirst-Of-Its-Kind Portable Germicidal Light Technology Disinfects High-Touch Clinical Surfaces in Seconds
Reducing healthcare-acquired infections (HAIs) remains a pressing issue within global healthcare systems. In the United States alone, 1.7 million patients contract HAIs annually, leading to approximately... Read more
Surgical Capacity Optimization Solution Helps Hospitals Boost OR Utilization
An innovative solution has the capability to transform surgical capacity utilization by targeting the root cause of surgical block time inefficiencies. Fujitsu Limited’s (Tokyo, Japan) Surgical Capacity... Read more
Game-Changing Innovation in Surgical Instrument Sterilization Significantly Improves OR Throughput
A groundbreaking innovation enables hospitals to significantly improve instrument processing time and throughput in operating rooms (ORs) and sterile processing departments. Turbett Surgical, Inc.... Read morePoint of Care
view channel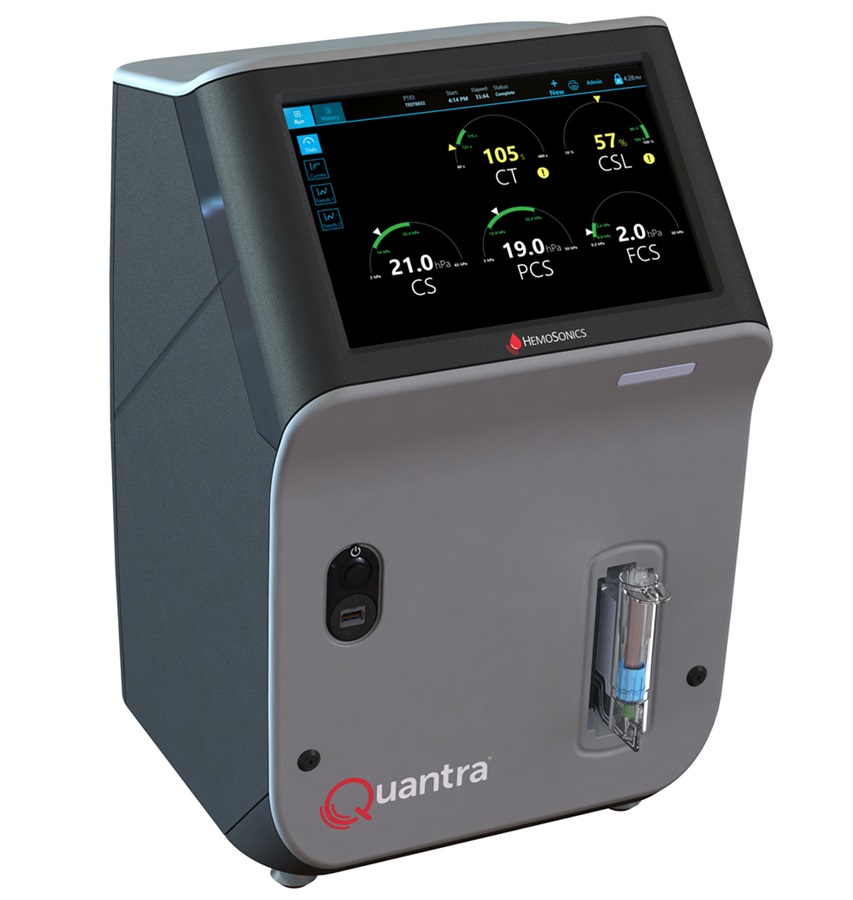
Critical Bleeding Management System to Help Hospitals Further Standardize Viscoelastic Testing
Surgical procedures are often accompanied by significant blood loss and the subsequent high likelihood of the need for allogeneic blood transfusions. These transfusions, while critical, are linked to various... Read more
Point of Care HIV Test Enables Early Infection Diagnosis for Infants
Early diagnosis and initiation of treatment are crucial for the survival of infants infected with HIV (human immunodeficiency virus). Without treatment, approximately 50% of infants who acquire HIV during... Read more
Whole Blood Rapid Test Aids Assessment of Concussion at Patient's Bedside
In the United States annually, approximately five million individuals seek emergency department care for traumatic brain injuries (TBIs), yet over half of those suspecting a concussion may never get it checked.... Read more
New Generation Glucose Hospital Meter System Ensures Accurate, Interference-Free and Safe Use
A new generation glucose hospital meter system now comes with several features that make hospital glucose testing easier and more secure while continuing to offer accuracy, freedom from interference, and... Read moreBusiness
view channel
Johnson & Johnson Acquires Cardiovascular Medical Device Company Shockwave Medical
Johnson & Johnson (New Brunswick, N.J., USA) and Shockwave Medical (Santa Clara, CA, USA) have entered into a definitive agreement under which Johnson & Johnson will acquire all of Shockwave’s... Read more