Rapid Sequencing Technique Accelerates Sepsis Diagnosis HCC
|
By HospiMedica International staff writers Posted on 16 Mar 2020 |
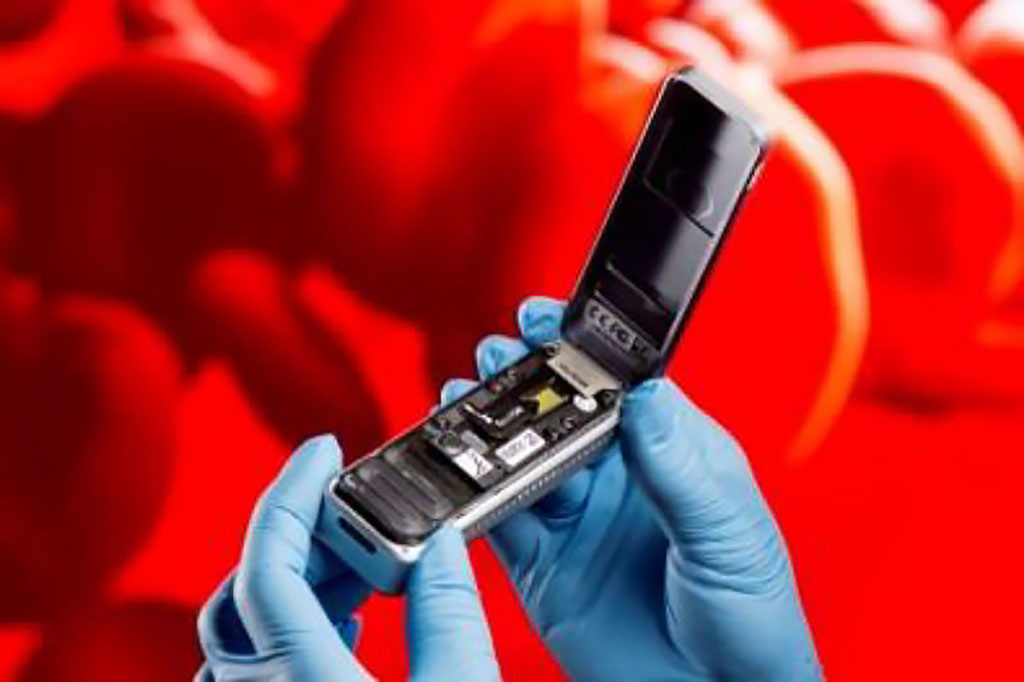

Image: The MinION handheld nanopore sequencer (Photo courtesy of IGB)
Real-time next-generation sequencing (NGS) allows accurate diagnosis of sepsis-causing agents within a few hours of drawing blood, claims a new study.
Researchers at the Fraunhofer Institute for Interfacial Engineering and Biotechnology (IGB; Stuttgart, Germany), Heidelberg University Hospital (Germany), and other institutions developed a diagnostic workflow that uses a handheld nanopore sequencer, the MinION, for real-time high-throughput sequencing of cell-free DNA from plasma, via nanopore sequencing of short fragments with low input amounts. The technique was used to analyze eight samples from four septic patients and three healthy controls, and subsequently validated against standard next-generation sequencing results.
The results revealed that with tailored bioinformatics workflows, all eight septic patient samples were found to be positive for relevant pathogens. When considering the time to diagnosis, the pathogens were identified within minutes; an extrapolation of real-time sequencing performance on a cohort of 239 septic patient samples revealed that with the MinION workflow, more than 90% of pathogen hits would have also been detected. The study was published in the March 2020 issue of The Journal of Molecular Diagnostics.
“With up to 50 million incident sepsis cases and 11 million sepsis-related deaths per year, sepsis represents a major cause of health loss,” said co-lead-investigator Thorsten Brenner, MD, of Heidelberg University Hospital. “Reliable and early identification of the pathogen enables rapid and the most appropriate antibiotic intervention, thereby increasing the chance of better outcomes and patient survival. Currently, standard-of-care diagnostics still rely on microbiological culturing of the respective pathogens, which in most cases do not provide timely positive results.”
The MinION, made by Oxford Nanopore Technologies (United Kingdom) is a pocket-sized device for real-time DNA and RNA sequencing, using up to 512 nanopore channels. Each consumable flow cell generates as much as 30 Gb of DNA sequence data or 7-12 million RNA reads. The MinION streams data into a PC or laptop using a high-speed USB 3.0 cable, or used alongside the MinIT device for real-time, powerful analyses.
Related Links:
Fraunhofer Institute for Interfacial Engineering and Biotechnology
Heidelberg University Hospital
Oxford Nanopore Technologies
Researchers at the Fraunhofer Institute for Interfacial Engineering and Biotechnology (IGB; Stuttgart, Germany), Heidelberg University Hospital (Germany), and other institutions developed a diagnostic workflow that uses a handheld nanopore sequencer, the MinION, for real-time high-throughput sequencing of cell-free DNA from plasma, via nanopore sequencing of short fragments with low input amounts. The technique was used to analyze eight samples from four septic patients and three healthy controls, and subsequently validated against standard next-generation sequencing results.
The results revealed that with tailored bioinformatics workflows, all eight septic patient samples were found to be positive for relevant pathogens. When considering the time to diagnosis, the pathogens were identified within minutes; an extrapolation of real-time sequencing performance on a cohort of 239 septic patient samples revealed that with the MinION workflow, more than 90% of pathogen hits would have also been detected. The study was published in the March 2020 issue of The Journal of Molecular Diagnostics.
“With up to 50 million incident sepsis cases and 11 million sepsis-related deaths per year, sepsis represents a major cause of health loss,” said co-lead-investigator Thorsten Brenner, MD, of Heidelberg University Hospital. “Reliable and early identification of the pathogen enables rapid and the most appropriate antibiotic intervention, thereby increasing the chance of better outcomes and patient survival. Currently, standard-of-care diagnostics still rely on microbiological culturing of the respective pathogens, which in most cases do not provide timely positive results.”
The MinION, made by Oxford Nanopore Technologies (United Kingdom) is a pocket-sized device for real-time DNA and RNA sequencing, using up to 512 nanopore channels. Each consumable flow cell generates as much as 30 Gb of DNA sequence data or 7-12 million RNA reads. The MinION streams data into a PC or laptop using a high-speed USB 3.0 cable, or used alongside the MinIT device for real-time, powerful analyses.
Related Links:
Fraunhofer Institute for Interfacial Engineering and Biotechnology
Heidelberg University Hospital
Oxford Nanopore Technologies
Latest Critical Care News
- Transcatheter Valve Replacement Outcomes Similar To Surgery, Finds Study
- Revascularization Improves Life Quality in Chronic Limb-Threatening Ischemia, Finds Study
- Powerful AI Risk Assessment Tool Predicts Outcomes in Heart Failure Patients
- Peptide-Based Hydrogels Repair Damaged Organs and Tissues On-The-Spot
- One-Hour Endoscopic Procedure Could Eliminate Need for Insulin for Type 2 Diabetes
- AI Can Prioritize Emergency Department Patients Requiring Urgent Treatment
- AI to Improve Diagnosis of Atrial Fibrillation
- Stretchable Microneedles to Help In Accurate Tracking of Abnormalities and Identifying Rapid Treatment
- Machine Learning Tool Identifies Rare, Undiagnosed Immune Disorders from Patient EHRs
- On-Skin Wearable Bioelectronic Device Paves Way for Intelligent Implants
- First-Of-Its-Kind Dissolvable Stent to Improve Outcomes for Patients with Severe PAD
- AI Brain-Age Estimation Technology Uses EEG Scans to Screen for Degenerative Diseases
- Wheeze-Counting Wearable Device Monitors Patient's Breathing In Real Time
- Wearable Multiplex Biosensors Could Revolutionize COPD Management
- New Low-Energy Defibrillation Method Controls Cardiac Arrhythmias
- New Machine Learning Models Help Predict Heart Disease Risk in Women
Channels
Artificial Intelligence
view channel
AI-Powered Algorithm to Revolutionize Detection of Atrial Fibrillation
Atrial fibrillation (AFib), a condition characterized by an irregular and often rapid heart rate, is linked to increased risks of stroke and heart failure. This is because the irregular heartbeat in AFib... Read more
AI Diagnostic Tool Accurately Detects Valvular Disorders Often Missed by Doctors
Doctors generally use stethoscopes to listen for the characteristic lub-dub sounds made by heart valves opening and closing. They also listen for less prominent sounds that indicate problems with these valves.... Read moreSurgical Techniques
view channel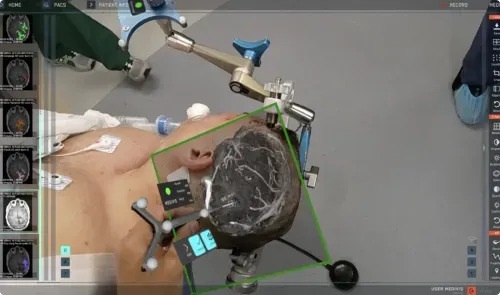
AR Surgical Technology Translates Complex 2D Medical Imaging to Enhance Accuracy
Surgeons often have to switch their focus between a patient’s data displayed on a screen or clipboard and the patient themselves during procedures. But that is about to change. Surgeons can now utilize... Read more
Miniaturized Snake-Like Probe Images Cerebral Arteries From Within
Endovascular interventions are being increasingly favored for treating strokes and cerebral artery diseases, but rely heavily on angiographical imaging that often struggles with limited contrast and spatial... Read morePatient Care
view channelFirst-Of-Its-Kind Portable Germicidal Light Technology Disinfects High-Touch Clinical Surfaces in Seconds
Reducing healthcare-acquired infections (HAIs) remains a pressing issue within global healthcare systems. In the United States alone, 1.7 million patients contract HAIs annually, leading to approximately... Read more
Surgical Capacity Optimization Solution Helps Hospitals Boost OR Utilization
An innovative solution has the capability to transform surgical capacity utilization by targeting the root cause of surgical block time inefficiencies. Fujitsu Limited’s (Tokyo, Japan) Surgical Capacity... Read more
Game-Changing Innovation in Surgical Instrument Sterilization Significantly Improves OR Throughput
A groundbreaking innovation enables hospitals to significantly improve instrument processing time and throughput in operating rooms (ORs) and sterile processing departments. Turbett Surgical, Inc.... Read moreHealth IT
view channel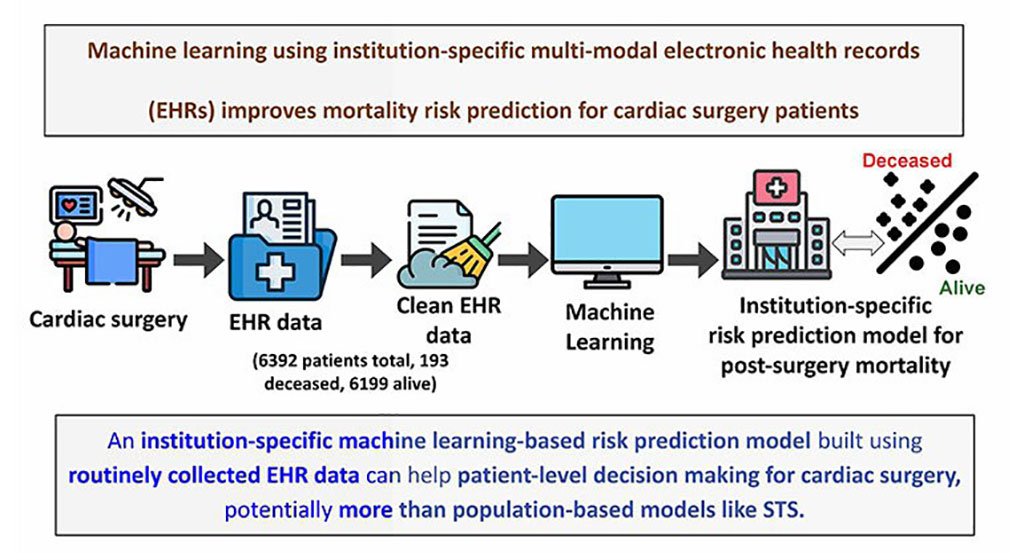
Machine Learning Model Improves Mortality Risk Prediction for Cardiac Surgery Patients
Machine learning algorithms have been deployed to create predictive models in various medical fields, with some demonstrating improved outcomes compared to their standard-of-care counterparts.... Read more
Strategic Collaboration to Develop and Integrate Generative AI into Healthcare
Top industry experts have underscored the immediate requirement for healthcare systems and hospitals to respond to severe cost and margin pressures. Close to half of U.S. hospitals ended 2022 in the red... Read more
AI-Enabled Operating Rooms Solution Helps Hospitals Maximize Utilization and Unlock Capacity
For healthcare organizations, optimizing operating room (OR) utilization during prime time hours is a complex challenge. Surgeons and clinics face difficulties in finding available slots for booking cases,... Read more
AI Predicts Pancreatic Cancer Three Years before Diagnosis from Patients’ Medical Records
Screening for common cancers like breast, cervix, and prostate cancer relies on relatively simple and highly effective techniques, such as mammograms, Pap smears, and blood tests. These methods have revolutionized... Read morePoint of Care
view channel
Critical Bleeding Management System to Help Hospitals Further Standardize Viscoelastic Testing
Surgical procedures are often accompanied by significant blood loss and the subsequent high likelihood of the need for allogeneic blood transfusions. These transfusions, while critical, are linked to various... Read more
Point of Care HIV Test Enables Early Infection Diagnosis for Infants
Early diagnosis and initiation of treatment are crucial for the survival of infants infected with HIV (human immunodeficiency virus). Without treatment, approximately 50% of infants who acquire HIV during... Read more
Whole Blood Rapid Test Aids Assessment of Concussion at Patient's Bedside
In the United States annually, approximately five million individuals seek emergency department care for traumatic brain injuries (TBIs), yet over half of those suspecting a concussion may never get it checked.... Read more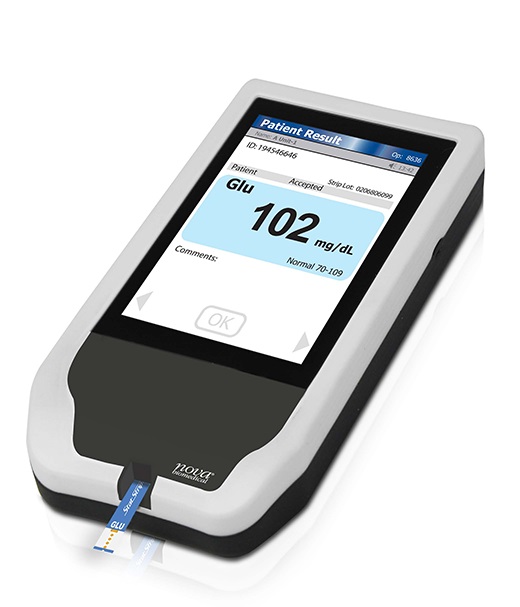
New Generation Glucose Hospital Meter System Ensures Accurate, Interference-Free and Safe Use
A new generation glucose hospital meter system now comes with several features that make hospital glucose testing easier and more secure while continuing to offer accuracy, freedom from interference, and... Read moreBusiness
view channel
Johnson & Johnson Acquires Cardiovascular Medical Device Company Shockwave Medical
Johnson & Johnson (New Brunswick, N.J., USA) and Shockwave Medical (Santa Clara, CA, USA) have entered into a definitive agreement under which Johnson & Johnson will acquire all of Shockwave’s... Read more