Mutation Could Allow SARS-CoV-2 to Develop Resistance to Gilead’s Remdesivir, Suggests New Research
|
By HospiMedica International staff writers Posted on 09 Oct 2020 |
Illustration
Researchers have identified how the Ebola virus and SARS-CoV-2 could mutate to develop resistance to treatment with Gilead Sciences’ (Foster City, CA, USA) investigational antiviral drug remdesivir.
Their study identified a single amino acid residue in the Ebola virus polymerase that conferred low-level resistance to remdesivir. More importantly, in addition to characterizing this particular mutation, the scientists related it to a resistance mutation observed in a similar structural motif of coronaviruses. Remdesivir is a nucleotide analog prodrug that has been clinically evaluated against both Ebola virus disease and COVID-19, and has recently received emergency use authorization (EUA) for the latter. Remdesivir was first characterized and evaluated as a potent inhibitor of the Ebola virus. The drug has also shown efficacy in mice and nonhuman primates against other highly pathogenic respiratory pathogens, including Nipah virus and both severe acute respiratory syndrome coronavirus (SARS-CoV-1) and Middle East respiratory syndrome coronavirus (MERS-CoV). Furthermore, preliminary evidence from clinical evaluations indicate that remdesivir shortens the recovery time of hospitalized COVID-19 patients presumably by blocking RNA replication of SARS-CoV-2. Remdesivir has been biochemically shown to inhibit the activity of Ebola virus large (L) RNA-dependent RNA polymerase (RdRp) as a non-obligate delayed chain terminator.
In line with the FDA’s recommendation for researchers to identify and characterize how viruses become resistant to drugs in order to gain a better understanding of their mechanism of action, scientists at the US Centers for Disease Control and Prevention (CDC) attempted to further understand the mechanism of Ebola virus inhibition and to identify determinants of resistance that may arise upon treatment of patients with remdesivir. Importantly, such determinants may naturally be present in other filoviruses, either known or yet to cross over into the human population. They found that remdesivir’s mechanism of action is common to both the Ebola virus as well as SARS-CoV-2.
The team serially passaged recombinant Ebola viruses with subclinical concentrations of remdesivir and demonstrated the reduced susceptibility of these viruses to remdesivir after 35 passages. The scientists identified a single-nucleotide variant (SNV) that emerged across six independent remdesivir-selected Ebola virus lineages; this mutation resulted in a non-conservative amino acid substitution at residue 548 (F548S) in the fingers sub-domain of the Ebola virus L RdRp. The scientists also examined this mutation in several contexts: a cell-based minigenome, a cell-free biochemical polymerase assay, as well as in a full-length infectious recombinant Ebola virus. In the context of the infectious virus, the F548S substitution recapitulated the reduced susceptibility phenotype to remdesivir, and potentially showed a marginal decrease in viral fitness compared to wild type. Thus, the study importantly identified a molecular marker for reduced remdesivir susceptibility.
The findings have implications for the surveillance of filovirus sequences, for treatment of Ebola virus patients with remdesivir or other similarly acting inhibitors, and for the development of future anti- Ebola virus therapies. Furthermore, comparative structural modeling of the Ebola virus and SARS-CoV-2 RdRp domains indicate remdesivir targets the polymerases of Ebola viruses and coronaviruses similarly, such that the findings may have implications for remdesivir treatment of COVID-19 patients.
Related Links:
Gilead Sciences
Their study identified a single amino acid residue in the Ebola virus polymerase that conferred low-level resistance to remdesivir. More importantly, in addition to characterizing this particular mutation, the scientists related it to a resistance mutation observed in a similar structural motif of coronaviruses. Remdesivir is a nucleotide analog prodrug that has been clinically evaluated against both Ebola virus disease and COVID-19, and has recently received emergency use authorization (EUA) for the latter. Remdesivir was first characterized and evaluated as a potent inhibitor of the Ebola virus. The drug has also shown efficacy in mice and nonhuman primates against other highly pathogenic respiratory pathogens, including Nipah virus and both severe acute respiratory syndrome coronavirus (SARS-CoV-1) and Middle East respiratory syndrome coronavirus (MERS-CoV). Furthermore, preliminary evidence from clinical evaluations indicate that remdesivir shortens the recovery time of hospitalized COVID-19 patients presumably by blocking RNA replication of SARS-CoV-2. Remdesivir has been biochemically shown to inhibit the activity of Ebola virus large (L) RNA-dependent RNA polymerase (RdRp) as a non-obligate delayed chain terminator.
In line with the FDA’s recommendation for researchers to identify and characterize how viruses become resistant to drugs in order to gain a better understanding of their mechanism of action, scientists at the US Centers for Disease Control and Prevention (CDC) attempted to further understand the mechanism of Ebola virus inhibition and to identify determinants of resistance that may arise upon treatment of patients with remdesivir. Importantly, such determinants may naturally be present in other filoviruses, either known or yet to cross over into the human population. They found that remdesivir’s mechanism of action is common to both the Ebola virus as well as SARS-CoV-2.
The team serially passaged recombinant Ebola viruses with subclinical concentrations of remdesivir and demonstrated the reduced susceptibility of these viruses to remdesivir after 35 passages. The scientists identified a single-nucleotide variant (SNV) that emerged across six independent remdesivir-selected Ebola virus lineages; this mutation resulted in a non-conservative amino acid substitution at residue 548 (F548S) in the fingers sub-domain of the Ebola virus L RdRp. The scientists also examined this mutation in several contexts: a cell-based minigenome, a cell-free biochemical polymerase assay, as well as in a full-length infectious recombinant Ebola virus. In the context of the infectious virus, the F548S substitution recapitulated the reduced susceptibility phenotype to remdesivir, and potentially showed a marginal decrease in viral fitness compared to wild type. Thus, the study importantly identified a molecular marker for reduced remdesivir susceptibility.
The findings have implications for the surveillance of filovirus sequences, for treatment of Ebola virus patients with remdesivir or other similarly acting inhibitors, and for the development of future anti- Ebola virus therapies. Furthermore, comparative structural modeling of the Ebola virus and SARS-CoV-2 RdRp domains indicate remdesivir targets the polymerases of Ebola viruses and coronaviruses similarly, such that the findings may have implications for remdesivir treatment of COVID-19 patients.
Related Links:
Gilead Sciences
Latest COVID-19 News
- Low-Cost System Detects SARS-CoV-2 Virus in Hospital Air Using High-Tech Bubbles
- World's First Inhalable COVID-19 Vaccine Approved in China
- COVID-19 Vaccine Patch Fights SARS-CoV-2 Variants Better than Needles
- Blood Viscosity Testing Can Predict Risk of Death in Hospitalized COVID-19 Patients
- ‘Covid Computer’ Uses AI to Detect COVID-19 from Chest CT Scans
- MRI Lung-Imaging Technique Shows Cause of Long-COVID Symptoms
- Chest CT Scans of COVID-19 Patients Could Help Distinguish Between SARS-CoV-2 Variants
- Specialized MRI Detects Lung Abnormalities in Non-Hospitalized Long COVID Patients
- AI Algorithm Identifies Hospitalized Patients at Highest Risk of Dying From COVID-19
- Sweat Sensor Detects Key Biomarkers That Provide Early Warning of COVID-19 and Flu
- Study Assesses Impact of COVID-19 on Ventilation/Perfusion Scintigraphy
- CT Imaging Study Finds Vaccination Reduces Risk of COVID-19 Associated Pulmonary Embolism
- Third Day in Hospital a ‘Tipping Point’ in Severity of COVID-19 Pneumonia
- Longer Interval Between COVID-19 Vaccines Generates Up to Nine Times as Many Antibodies
- AI Model for Monitoring COVID-19 Predicts Mortality Within First 30 Days of Admission
- AI Predicts COVID Prognosis at Near-Expert Level Based Off CT Scans
Channels
Artificial Intelligence
view channel
AI-Powered Algorithm to Revolutionize Detection of Atrial Fibrillation
Atrial fibrillation (AFib), a condition characterized by an irregular and often rapid heart rate, is linked to increased risks of stroke and heart failure. This is because the irregular heartbeat in AFib... Read more
AI Diagnostic Tool Accurately Detects Valvular Disorders Often Missed by Doctors
Doctors generally use stethoscopes to listen for the characteristic lub-dub sounds made by heart valves opening and closing. They also listen for less prominent sounds that indicate problems with these valves.... Read moreCritical Care
view channel
Powerful AI Risk Assessment Tool Predicts Outcomes in Heart Failure Patients
Heart failure is a serious condition where the heart cannot pump sufficient blood to meet the body's needs, leading to symptoms like fatigue, weakness, and swelling in the legs and feet, and it can ultimately... Read more
Peptide-Based Hydrogels Repair Damaged Organs and Tissues On-The-Spot
Scientists have ingeniously combined biomedical expertise with nature-inspired engineering to develop a jelly-like material that holds significant promise for immediate repairs to a wide variety of damaged... Read more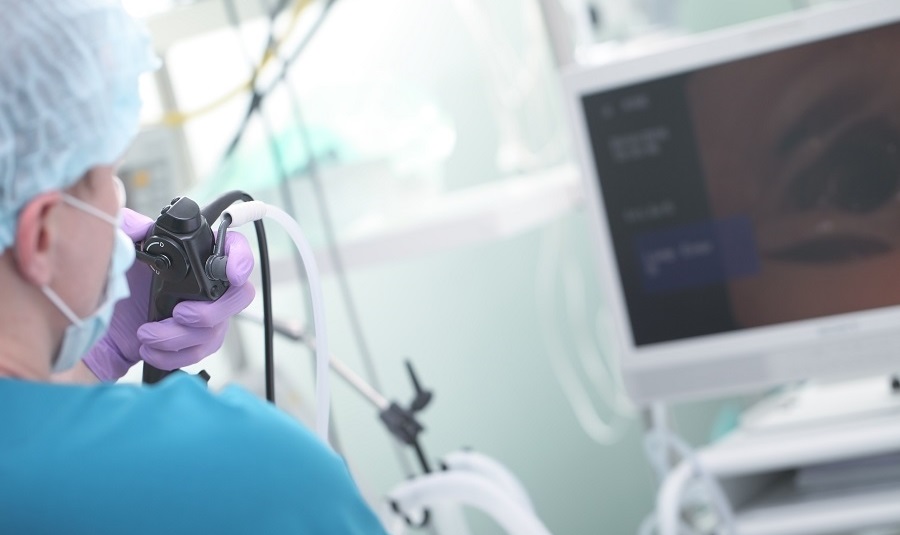
One-Hour Endoscopic Procedure Could Eliminate Need for Insulin for Type 2 Diabetes
Over 37 million Americans are diagnosed with diabetes, and more than 90% of these cases are Type 2 diabetes. This form of diabetes is most commonly seen in individuals over 45, though an increasing number... Read moreSurgical Techniques
view channel
Miniaturized Implantable Multi-Sensors Device to Monitor Vessels Health
Researchers have embarked on a project to develop a multi-sensing device that can be implanted into blood vessels like peripheral veins or arteries to monitor a range of bodily parameters and overall health status.... Read more
Tiny Robots Made Out Of Carbon Could Conduct Colonoscopy, Pelvic Exam or Blood Test
Researchers at the University of Alberta (Edmonton, AB, Canada) are developing cutting-edge robots so tiny that they are invisible to the naked eye but are capable of traveling through the human body to... Read more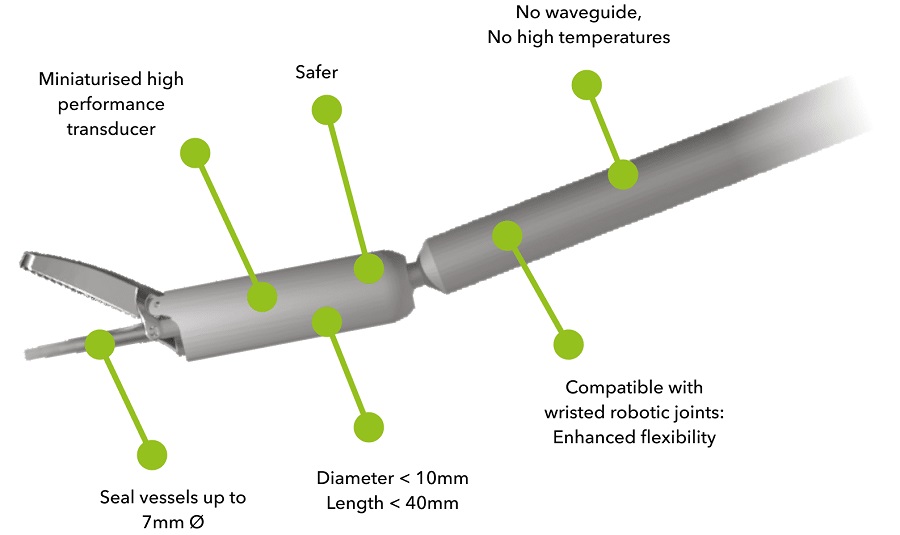
Miniaturized Ultrasonic Scalpel Enables Faster and Safer Robotic-Assisted Surgery
Robot-assisted surgery (RAS) has gained significant popularity in recent years and is now extensively used across various surgical fields such as urology, gynecology, and cardiology. These surgeries, performed... Read morePatient Care
view channelFirst-Of-Its-Kind Portable Germicidal Light Technology Disinfects High-Touch Clinical Surfaces in Seconds
Reducing healthcare-acquired infections (HAIs) remains a pressing issue within global healthcare systems. In the United States alone, 1.7 million patients contract HAIs annually, leading to approximately... Read more
Surgical Capacity Optimization Solution Helps Hospitals Boost OR Utilization
An innovative solution has the capability to transform surgical capacity utilization by targeting the root cause of surgical block time inefficiencies. Fujitsu Limited’s (Tokyo, Japan) Surgical Capacity... Read more
Game-Changing Innovation in Surgical Instrument Sterilization Significantly Improves OR Throughput
A groundbreaking innovation enables hospitals to significantly improve instrument processing time and throughput in operating rooms (ORs) and sterile processing departments. Turbett Surgical, Inc.... Read moreHealth IT
view channel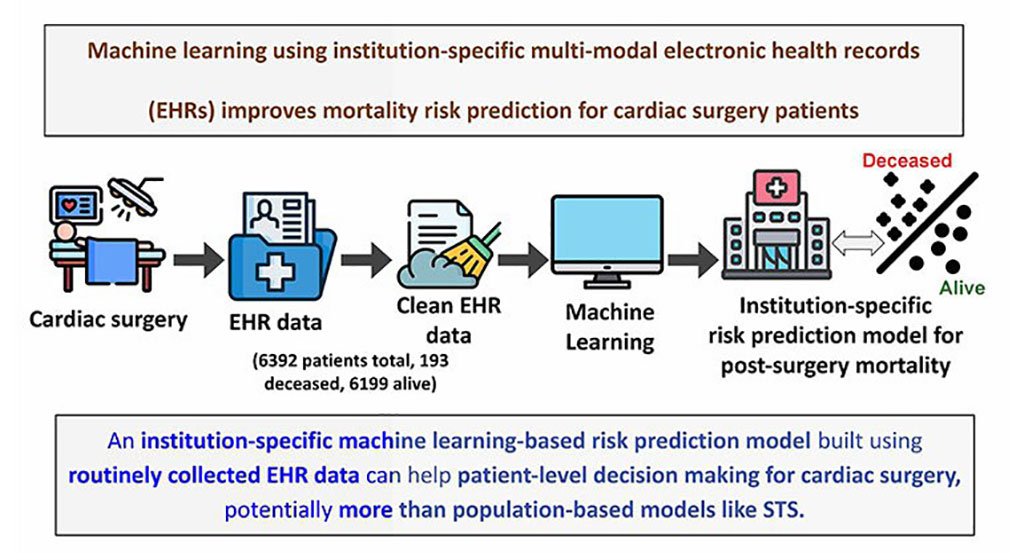
Machine Learning Model Improves Mortality Risk Prediction for Cardiac Surgery Patients
Machine learning algorithms have been deployed to create predictive models in various medical fields, with some demonstrating improved outcomes compared to their standard-of-care counterparts.... Read more
Strategic Collaboration to Develop and Integrate Generative AI into Healthcare
Top industry experts have underscored the immediate requirement for healthcare systems and hospitals to respond to severe cost and margin pressures. Close to half of U.S. hospitals ended 2022 in the red... Read more
AI-Enabled Operating Rooms Solution Helps Hospitals Maximize Utilization and Unlock Capacity
For healthcare organizations, optimizing operating room (OR) utilization during prime time hours is a complex challenge. Surgeons and clinics face difficulties in finding available slots for booking cases,... Read more
AI Predicts Pancreatic Cancer Three Years before Diagnosis from Patients’ Medical Records
Screening for common cancers like breast, cervix, and prostate cancer relies on relatively simple and highly effective techniques, such as mammograms, Pap smears, and blood tests. These methods have revolutionized... Read morePoint of Care
view channel
Critical Bleeding Management System to Help Hospitals Further Standardize Viscoelastic Testing
Surgical procedures are often accompanied by significant blood loss and the subsequent high likelihood of the need for allogeneic blood transfusions. These transfusions, while critical, are linked to various... Read more
Point of Care HIV Test Enables Early Infection Diagnosis for Infants
Early diagnosis and initiation of treatment are crucial for the survival of infants infected with HIV (human immunodeficiency virus). Without treatment, approximately 50% of infants who acquire HIV during... Read more
Whole Blood Rapid Test Aids Assessment of Concussion at Patient's Bedside
In the United States annually, approximately five million individuals seek emergency department care for traumatic brain injuries (TBIs), yet over half of those suspecting a concussion may never get it checked.... Read more
New Generation Glucose Hospital Meter System Ensures Accurate, Interference-Free and Safe Use
A new generation glucose hospital meter system now comes with several features that make hospital glucose testing easier and more secure while continuing to offer accuracy, freedom from interference, and... Read moreBusiness
view channel
Johnson & Johnson Acquires Cardiovascular Medical Device Company Shockwave Medical
Johnson & Johnson (New Brunswick, N.J., USA) and Shockwave Medical (Santa Clara, CA, USA) have entered into a definitive agreement under which Johnson & Johnson will acquire all of Shockwave’s... Read more