New Artificial Intelligence Method Helps Design Better COVID-19 Antibody Drugs
|
By HospiMedica International staff writers Posted on 20 Apr 2021 |

Illustration
Machine learning methods can help to optimize the development of COVID-19 antibody drugs, leading to active substances with improved properties, also with regard to tolerability in the body, according to researchers.
Scientists at ETH Zürich (Zürich, Switzerland) have developed a machine learning method that supports the optimization phase, helping to develop more effective antibody drugs. Antibodies are not only produced by our immune cells to fight viruses and other pathogens in the body. For a few decades now, medicine has also been using antibodies produced by biotechnology as drugs. This is because antibodies are extremely good at binding specifically to molecular structures according to the lock-and-key principle. Their use ranges from oncology to the treatment of autoimmune diseases and neurodegenerative conditions.
However, developing such antibody drugs is anything but simple. The basic requirement is for an antibody to bind to its target molecule in an optimal way. At the same time, an antibody drug must fulfill a host of additional criteria. For example, it should not trigger an immune response in the body, it should be efficient to produce using biotechnology, and it should remain stable over a long period of time. Once scientists have found an antibody that binds to the desired molecular target structure, the development process is far from over. Rather, this marks the start of a phase in which researchers use bioengineering to try to improve the antibody’s properties.
When researchers optimize an entire antibody molecule in its therapeutic form (i.e. not just a fragment of an antibody), it used to start with an antibody lead candidate that binds reasonably well to the desired target structure. Then researchers randomly mutate the gene that carries the blueprint for the antibody in order to produce a few thousand related antibody candidates in the lab. The next step is to search among them to find the ones that bind best to the target structure. The ETH researchers are now using machine learning to increase the initial set of antibodies to be tested to several million.
The researchers provided the proof of concept for their new method using Roche’s antibody cancer drug Herceptin, which has been on the market for 20 years. Starting out from the DNA sequence of the Herceptin antibody, the ETH researchers created about 40,000 related antibodies using a CRISPR mutation method they developed a few years ago. Experiments showed that 10,000 of them bound well to the target protein in question, a specific cell surface protein. The scientists used the DNA sequences of these 40,000 antibodies to train a machine learning algorithm. They then applied the trained algorithm to search a database of 70 million potential antibody DNA sequences. For these 70 million candidates, the algorithm predicted how well the corresponding antibodies would bind to the target protein, resulting in a list of millions of sequences expected to bind.
Using further computer models, the scientists predicted how well these millions of sequences would meet the additional criteria for drug development (tolerance, production, physical properties). This reduced the number of candidate sequences to 8,000. From the list of optimized candidate sequences on their computer, the scientists selected 55 sequences from which to produce antibodies in the lab and characterize their properties. Subsequent experiments showed that several of them bound even better to the target protein than Herceptin itself, as well as being easier to produce and more stable than Herceptin. The ETH scientists are now applying their AI method to optimize antibody drugs that are in clinical development.
“With automated processes, you can test a few thousand therapeutic candidates in a lab. But it is not really feasible to screen any more than that,” said Sai Reddy, a professor at the Department of Biosystems Science and Engineering at ETH Zurich who led the study. “Typically, the best dozen antibodies from this screening move on to the next step and are tested for how well they meet additional criteria. “Ultimately, this approach lets you identify the best antibody from a group of a few thousand.”
Related Links:
ETH Zürich
Scientists at ETH Zürich (Zürich, Switzerland) have developed a machine learning method that supports the optimization phase, helping to develop more effective antibody drugs. Antibodies are not only produced by our immune cells to fight viruses and other pathogens in the body. For a few decades now, medicine has also been using antibodies produced by biotechnology as drugs. This is because antibodies are extremely good at binding specifically to molecular structures according to the lock-and-key principle. Their use ranges from oncology to the treatment of autoimmune diseases and neurodegenerative conditions.
However, developing such antibody drugs is anything but simple. The basic requirement is for an antibody to bind to its target molecule in an optimal way. At the same time, an antibody drug must fulfill a host of additional criteria. For example, it should not trigger an immune response in the body, it should be efficient to produce using biotechnology, and it should remain stable over a long period of time. Once scientists have found an antibody that binds to the desired molecular target structure, the development process is far from over. Rather, this marks the start of a phase in which researchers use bioengineering to try to improve the antibody’s properties.
When researchers optimize an entire antibody molecule in its therapeutic form (i.e. not just a fragment of an antibody), it used to start with an antibody lead candidate that binds reasonably well to the desired target structure. Then researchers randomly mutate the gene that carries the blueprint for the antibody in order to produce a few thousand related antibody candidates in the lab. The next step is to search among them to find the ones that bind best to the target structure. The ETH researchers are now using machine learning to increase the initial set of antibodies to be tested to several million.
The researchers provided the proof of concept for their new method using Roche’s antibody cancer drug Herceptin, which has been on the market for 20 years. Starting out from the DNA sequence of the Herceptin antibody, the ETH researchers created about 40,000 related antibodies using a CRISPR mutation method they developed a few years ago. Experiments showed that 10,000 of them bound well to the target protein in question, a specific cell surface protein. The scientists used the DNA sequences of these 40,000 antibodies to train a machine learning algorithm. They then applied the trained algorithm to search a database of 70 million potential antibody DNA sequences. For these 70 million candidates, the algorithm predicted how well the corresponding antibodies would bind to the target protein, resulting in a list of millions of sequences expected to bind.
Using further computer models, the scientists predicted how well these millions of sequences would meet the additional criteria for drug development (tolerance, production, physical properties). This reduced the number of candidate sequences to 8,000. From the list of optimized candidate sequences on their computer, the scientists selected 55 sequences from which to produce antibodies in the lab and characterize their properties. Subsequent experiments showed that several of them bound even better to the target protein than Herceptin itself, as well as being easier to produce and more stable than Herceptin. The ETH scientists are now applying their AI method to optimize antibody drugs that are in clinical development.
“With automated processes, you can test a few thousand therapeutic candidates in a lab. But it is not really feasible to screen any more than that,” said Sai Reddy, a professor at the Department of Biosystems Science and Engineering at ETH Zurich who led the study. “Typically, the best dozen antibodies from this screening move on to the next step and are tested for how well they meet additional criteria. “Ultimately, this approach lets you identify the best antibody from a group of a few thousand.”
Related Links:
ETH Zürich
Latest COVID-19 News
- Low-Cost System Detects SARS-CoV-2 Virus in Hospital Air Using High-Tech Bubbles
- World's First Inhalable COVID-19 Vaccine Approved in China
- COVID-19 Vaccine Patch Fights SARS-CoV-2 Variants Better than Needles
- Blood Viscosity Testing Can Predict Risk of Death in Hospitalized COVID-19 Patients
- ‘Covid Computer’ Uses AI to Detect COVID-19 from Chest CT Scans
- MRI Lung-Imaging Technique Shows Cause of Long-COVID Symptoms
- Chest CT Scans of COVID-19 Patients Could Help Distinguish Between SARS-CoV-2 Variants
- Specialized MRI Detects Lung Abnormalities in Non-Hospitalized Long COVID Patients
- AI Algorithm Identifies Hospitalized Patients at Highest Risk of Dying From COVID-19
- Sweat Sensor Detects Key Biomarkers That Provide Early Warning of COVID-19 and Flu
- Study Assesses Impact of COVID-19 on Ventilation/Perfusion Scintigraphy
- CT Imaging Study Finds Vaccination Reduces Risk of COVID-19 Associated Pulmonary Embolism
- Third Day in Hospital a ‘Tipping Point’ in Severity of COVID-19 Pneumonia
- Longer Interval Between COVID-19 Vaccines Generates Up to Nine Times as Many Antibodies
- AI Model for Monitoring COVID-19 Predicts Mortality Within First 30 Days of Admission
- AI Predicts COVID Prognosis at Near-Expert Level Based Off CT Scans
Channels
Artificial Intelligence
view channel
AI-Powered Algorithm to Revolutionize Detection of Atrial Fibrillation
Atrial fibrillation (AFib), a condition characterized by an irregular and often rapid heart rate, is linked to increased risks of stroke and heart failure. This is because the irregular heartbeat in AFib... Read more
AI Diagnostic Tool Accurately Detects Valvular Disorders Often Missed by Doctors
Doctors generally use stethoscopes to listen for the characteristic lub-dub sounds made by heart valves opening and closing. They also listen for less prominent sounds that indicate problems with these valves.... Read moreCritical Care
view channel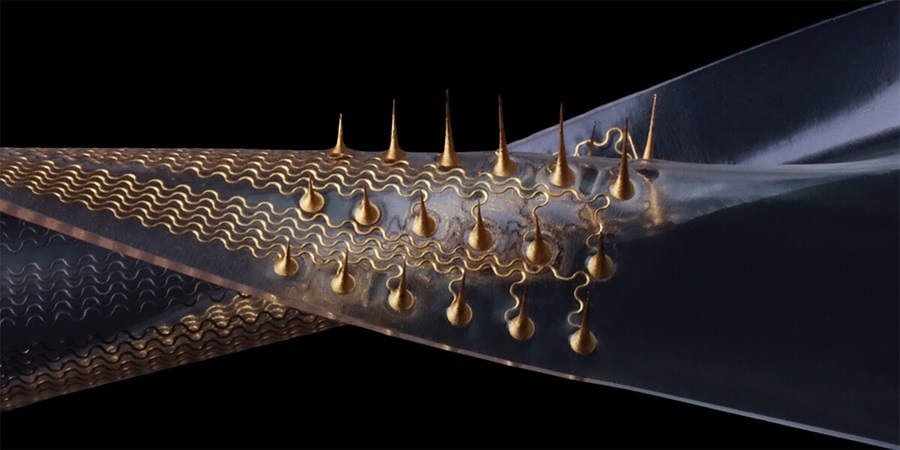
Stretchable Microneedles to Help In Accurate Tracking of Abnormalities and Identifying Rapid Treatment
The field of personalized medicine is transforming rapidly, with advancements like wearable devices and home testing kits making it increasingly easy to monitor a wide range of health metrics, from heart... Read more
Machine Learning Tool Identifies Rare, Undiagnosed Immune Disorders from Patient EHRs
Patients suffering from rare diseases often endure extensive delays in receiving accurate diagnoses and treatments, which can lead to unnecessary tests, worsening health, psychological strain, and significant... Read more
On-Skin Wearable Bioelectronic Device Paves Way for Intelligent Implants
A team of researchers at the University of Missouri (Columbia, MO, USA) has achieved a milestone in developing a state-of-the-art on-skin wearable bioelectronic device. This development comes from a lab... Read more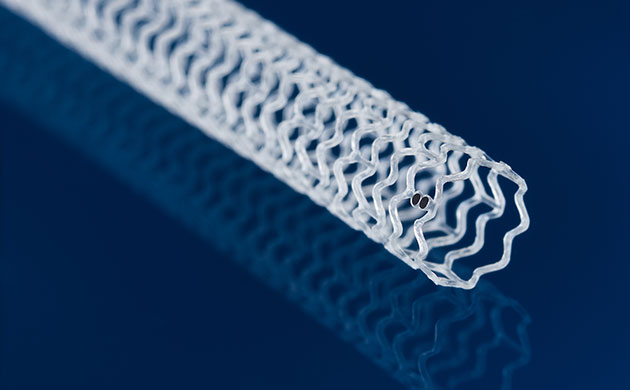
First-Of-Its-Kind Dissolvable Stent to Improve Outcomes for Patients with Severe PAD
Peripheral artery disease (PAD) affects millions and presents serious health risks, particularly its severe form, chronic limb-threatening ischemia (CLTI). CLTI develops when arteries are blocked by plaque,... Read moreSurgical Techniques
view channel
World’s Smallest Laser Probe for Brain Procedures Facilitates Ablation of Full Range of Targets
A new probe enhances the ablation capabilities for a broad spectrum of oncology and epilepsy targets, including pediatric applications, by incorporating advanced laser and cooling technologies to support... Read more.jpg)
Artificial Intelligence Broadens Diagnostic Abilities of Conventional Coronary Angiography
Coronary angiography is an essential diagnostic tool used globally to identify coronary artery disease (CAD), with millions of procedures conducted annually. Traditionally, data from coronary angiograms... Read moreAI-Powered Surgical Visualization Tool Supports Surgeons' Visual Recognition in Real Time
Connective tissue serves as an essential landmark in surgical navigation, often referred to as the "dissection plane" or "holy plane." Its accurate identification is vital for achieving safe and effective... Read moreHandheld Device for Fluorescence-Guided Surgery a Game Changer for Removal of High-Grade Glioma Brain Tumors
Grade III or IV gliomas are among the most common and deadly brain tumors, with around 20,000 cases annually in the U.S. and 1.2 million globally. These tumors are very aggressive and tend to infiltrate... Read morePatient Care
view channelFirst-Of-Its-Kind Portable Germicidal Light Technology Disinfects High-Touch Clinical Surfaces in Seconds
Reducing healthcare-acquired infections (HAIs) remains a pressing issue within global healthcare systems. In the United States alone, 1.7 million patients contract HAIs annually, leading to approximately... Read more
Surgical Capacity Optimization Solution Helps Hospitals Boost OR Utilization
An innovative solution has the capability to transform surgical capacity utilization by targeting the root cause of surgical block time inefficiencies. Fujitsu Limited’s (Tokyo, Japan) Surgical Capacity... Read more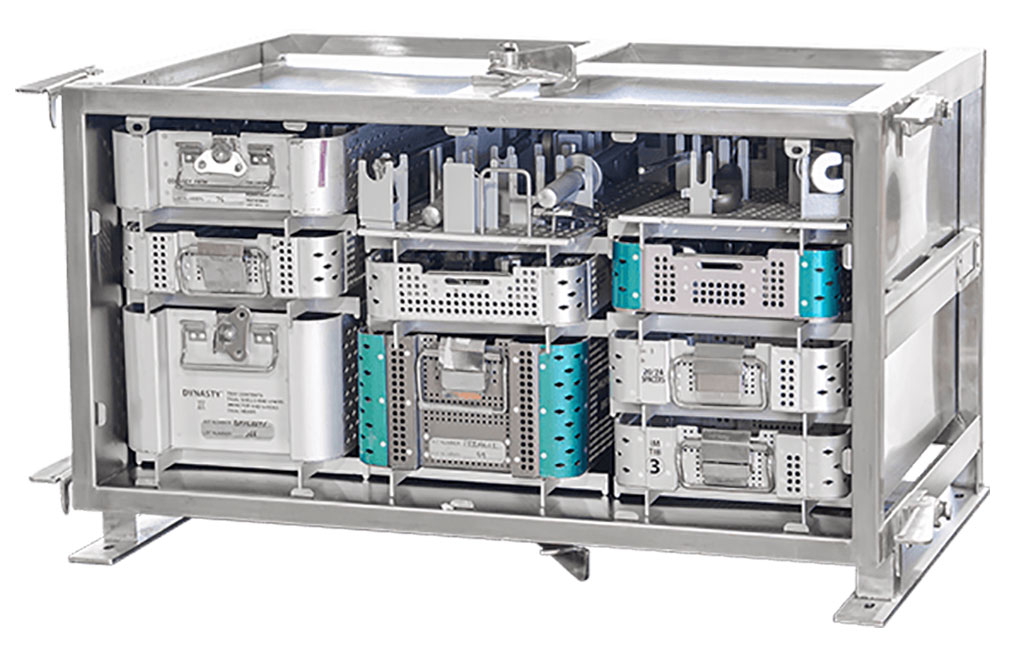
Game-Changing Innovation in Surgical Instrument Sterilization Significantly Improves OR Throughput
A groundbreaking innovation enables hospitals to significantly improve instrument processing time and throughput in operating rooms (ORs) and sterile processing departments. Turbett Surgical, Inc.... Read moreHealth IT
view channel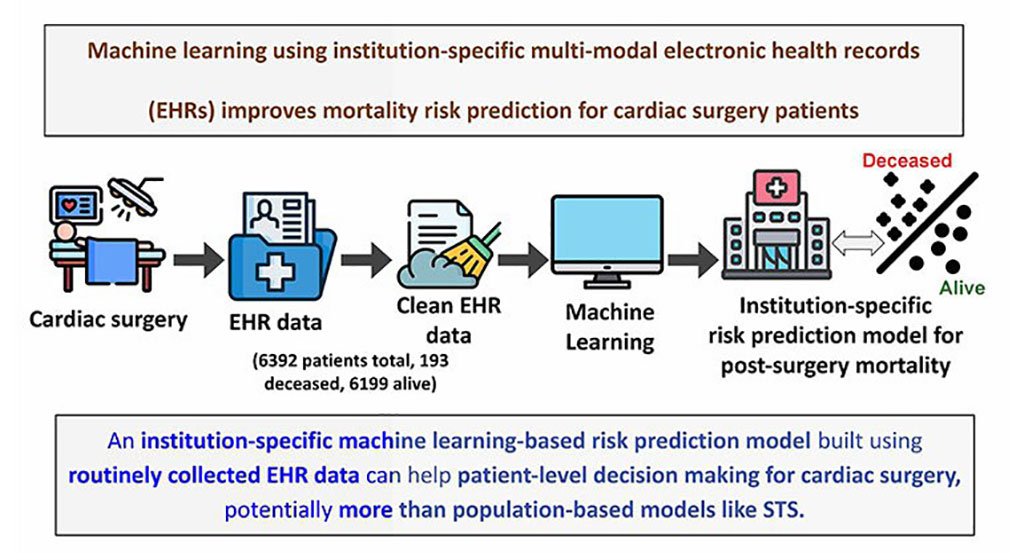
Machine Learning Model Improves Mortality Risk Prediction for Cardiac Surgery Patients
Machine learning algorithms have been deployed to create predictive models in various medical fields, with some demonstrating improved outcomes compared to their standard-of-care counterparts.... Read more
Strategic Collaboration to Develop and Integrate Generative AI into Healthcare
Top industry experts have underscored the immediate requirement for healthcare systems and hospitals to respond to severe cost and margin pressures. Close to half of U.S. hospitals ended 2022 in the red... Read more
AI-Enabled Operating Rooms Solution Helps Hospitals Maximize Utilization and Unlock Capacity
For healthcare organizations, optimizing operating room (OR) utilization during prime time hours is a complex challenge. Surgeons and clinics face difficulties in finding available slots for booking cases,... Read more
AI Predicts Pancreatic Cancer Three Years before Diagnosis from Patients’ Medical Records
Screening for common cancers like breast, cervix, and prostate cancer relies on relatively simple and highly effective techniques, such as mammograms, Pap smears, and blood tests. These methods have revolutionized... Read morePoint of Care
view channel
Critical Bleeding Management System to Help Hospitals Further Standardize Viscoelastic Testing
Surgical procedures are often accompanied by significant blood loss and the subsequent high likelihood of the need for allogeneic blood transfusions. These transfusions, while critical, are linked to various... Read more
Point of Care HIV Test Enables Early Infection Diagnosis for Infants
Early diagnosis and initiation of treatment are crucial for the survival of infants infected with HIV (human immunodeficiency virus). Without treatment, approximately 50% of infants who acquire HIV during... Read more
Whole Blood Rapid Test Aids Assessment of Concussion at Patient's Bedside
In the United States annually, approximately five million individuals seek emergency department care for traumatic brain injuries (TBIs), yet over half of those suspecting a concussion may never get it checked.... Read more
New Generation Glucose Hospital Meter System Ensures Accurate, Interference-Free and Safe Use
A new generation glucose hospital meter system now comes with several features that make hospital glucose testing easier and more secure while continuing to offer accuracy, freedom from interference, and... Read moreBusiness
view channel
Johnson & Johnson Acquires Cardiovascular Medical Device Company Shockwave Medical
Johnson & Johnson (New Brunswick, N.J., USA) and Shockwave Medical (Santa Clara, CA, USA) have entered into a definitive agreement under which Johnson & Johnson will acquire all of Shockwave’s... Read more