RNA Structures of SARS-CoV-2 Reveal Potential Drug Targets
|
By HospiMedica International staff writers Posted on 12 Nov 2020 |
Illustration
Researchers who studied the SARS-CoV-2 coronavirus RNA genome structure in detail have identified potential targets for the development of drugs against the virus.
COVID-19 is caused by SARS-CoV-2, a betacoronavirus with a linear single-stranded, positive-sense RNA genome. Similar to those in other RNA viruses, the SARS-CoV-2 RNA structures are expected to play a crucial role in how the coronavirus replicates in human cells. Despite this importance, only a handful of functionally relevant coronavirus structural RNA elements have been studied to date. Therefore, researchers from the University of Groningen (Groningen, Netherlands), together with scientists from the the International Institute of Molecular and Cell Biology (Warsaw, Poland) and Leiden University (Leiden, Netherlands), performed an extensive characterization of the SARS-CoV-2 RNA genome structure using various advanced techniques.
The study involved RNA structure probing to obtain single-base resolution secondary structure maps of the full SARS-CoV-2 coronavirus genome both in vitro and in living infected cells. Subsequently, the team identified at least 87 regions in the SARS-CoV-2 RNA sequence that appears to form well-defined compact structures. Of these, at least 10% are under strong evolutionary selection pressure among coronaviruses, suggesting functional relevance. Importantly, this is the first time that the structure of the entire coronavirus RNA (one of the longest viral RNAs with approximately 30,000 nucleotides) was determined.
Also, pockets were identified in some RNA structures that could be targeted by small molecules to hamper the function of the viral RNA. The scientists also identified parts of the SARS-CoV-2 RNA that are intrinsically unstructured. Adding short nucleic acid strands that can bind to these viral RNA sections would create double-stranded regions, which are naturally targeted by enzymes inside human cells. Thus, the collaborative research has established a firm foundation for future work aimed at developing potential small-molecule drugs for the treatment of SARS-CoV-2 infections and possibly also infections by other coronaviruses.
“We first identified the structures in vitro, and subsequently confirmed their presence in the RNA of viruses inside cells,” said Dr. Danny Incarnato from the University of Groningen who coordinated the study. “This means that our results are very robust. Furthermore, a number of the structures are conserved between different coronaviruses, meaning that a successful drug targeting SARS-CoV-2 could also be effective against future new virus strains.”
Related Links:
University of Groningen
International Institute of Molecular and Cell Biology
Leiden University
COVID-19 is caused by SARS-CoV-2, a betacoronavirus with a linear single-stranded, positive-sense RNA genome. Similar to those in other RNA viruses, the SARS-CoV-2 RNA structures are expected to play a crucial role in how the coronavirus replicates in human cells. Despite this importance, only a handful of functionally relevant coronavirus structural RNA elements have been studied to date. Therefore, researchers from the University of Groningen (Groningen, Netherlands), together with scientists from the the International Institute of Molecular and Cell Biology (Warsaw, Poland) and Leiden University (Leiden, Netherlands), performed an extensive characterization of the SARS-CoV-2 RNA genome structure using various advanced techniques.
The study involved RNA structure probing to obtain single-base resolution secondary structure maps of the full SARS-CoV-2 coronavirus genome both in vitro and in living infected cells. Subsequently, the team identified at least 87 regions in the SARS-CoV-2 RNA sequence that appears to form well-defined compact structures. Of these, at least 10% are under strong evolutionary selection pressure among coronaviruses, suggesting functional relevance. Importantly, this is the first time that the structure of the entire coronavirus RNA (one of the longest viral RNAs with approximately 30,000 nucleotides) was determined.
Also, pockets were identified in some RNA structures that could be targeted by small molecules to hamper the function of the viral RNA. The scientists also identified parts of the SARS-CoV-2 RNA that are intrinsically unstructured. Adding short nucleic acid strands that can bind to these viral RNA sections would create double-stranded regions, which are naturally targeted by enzymes inside human cells. Thus, the collaborative research has established a firm foundation for future work aimed at developing potential small-molecule drugs for the treatment of SARS-CoV-2 infections and possibly also infections by other coronaviruses.
“We first identified the structures in vitro, and subsequently confirmed their presence in the RNA of viruses inside cells,” said Dr. Danny Incarnato from the University of Groningen who coordinated the study. “This means that our results are very robust. Furthermore, a number of the structures are conserved between different coronaviruses, meaning that a successful drug targeting SARS-CoV-2 could also be effective against future new virus strains.”
Related Links:
University of Groningen
International Institute of Molecular and Cell Biology
Leiden University
Latest COVID-19 News
- Low-Cost System Detects SARS-CoV-2 Virus in Hospital Air Using High-Tech Bubbles
- World's First Inhalable COVID-19 Vaccine Approved in China
- COVID-19 Vaccine Patch Fights SARS-CoV-2 Variants Better than Needles
- Blood Viscosity Testing Can Predict Risk of Death in Hospitalized COVID-19 Patients
- ‘Covid Computer’ Uses AI to Detect COVID-19 from Chest CT Scans
- MRI Lung-Imaging Technique Shows Cause of Long-COVID Symptoms
- Chest CT Scans of COVID-19 Patients Could Help Distinguish Between SARS-CoV-2 Variants
- Specialized MRI Detects Lung Abnormalities in Non-Hospitalized Long COVID Patients
- AI Algorithm Identifies Hospitalized Patients at Highest Risk of Dying From COVID-19
- Sweat Sensor Detects Key Biomarkers That Provide Early Warning of COVID-19 and Flu
- Study Assesses Impact of COVID-19 on Ventilation/Perfusion Scintigraphy
- CT Imaging Study Finds Vaccination Reduces Risk of COVID-19 Associated Pulmonary Embolism
- Third Day in Hospital a ‘Tipping Point’ in Severity of COVID-19 Pneumonia
- Longer Interval Between COVID-19 Vaccines Generates Up to Nine Times as Many Antibodies
- AI Model for Monitoring COVID-19 Predicts Mortality Within First 30 Days of Admission
- AI Predicts COVID Prognosis at Near-Expert Level Based Off CT Scans
Channels
Artificial Intelligence
view channel
AI-Powered Algorithm to Revolutionize Detection of Atrial Fibrillation
Atrial fibrillation (AFib), a condition characterized by an irregular and often rapid heart rate, is linked to increased risks of stroke and heart failure. This is because the irregular heartbeat in AFib... Read more
AI Diagnostic Tool Accurately Detects Valvular Disorders Often Missed by Doctors
Doctors generally use stethoscopes to listen for the characteristic lub-dub sounds made by heart valves opening and closing. They also listen for less prominent sounds that indicate problems with these valves.... Read moreCritical Care
view channel
AI Brain-Age Estimation Technology Uses EEG Scans to Screen for Degenerative Diseases
As individuals age, so do their brains. Premature aging of the brain can lead to age-related conditions such as mild cognitive impairment, dementia, or Parkinson's disease. The ability to determine "brain... Read more
Wheeze-Counting Wearable Device Monitors Patient's Breathing In Real Time
Lung diseases like asthma, chronic obstructive pulmonary disease (COPD), lung cancer, bronchitis, and infections such as pneumonia, rank among the leading causes of death worldwide. Traditionally, medical... Read moreSurgical Techniques
view channel
Robotic Nerve ‘Cuffs’ Could Treat Various Neurological Conditions
Electric nerve implants serve dual functions: they can either stimulate or block signals in specific nerves. For example, they may alleviate pain by inhibiting pain signals or restore movement in paralyzed... Read more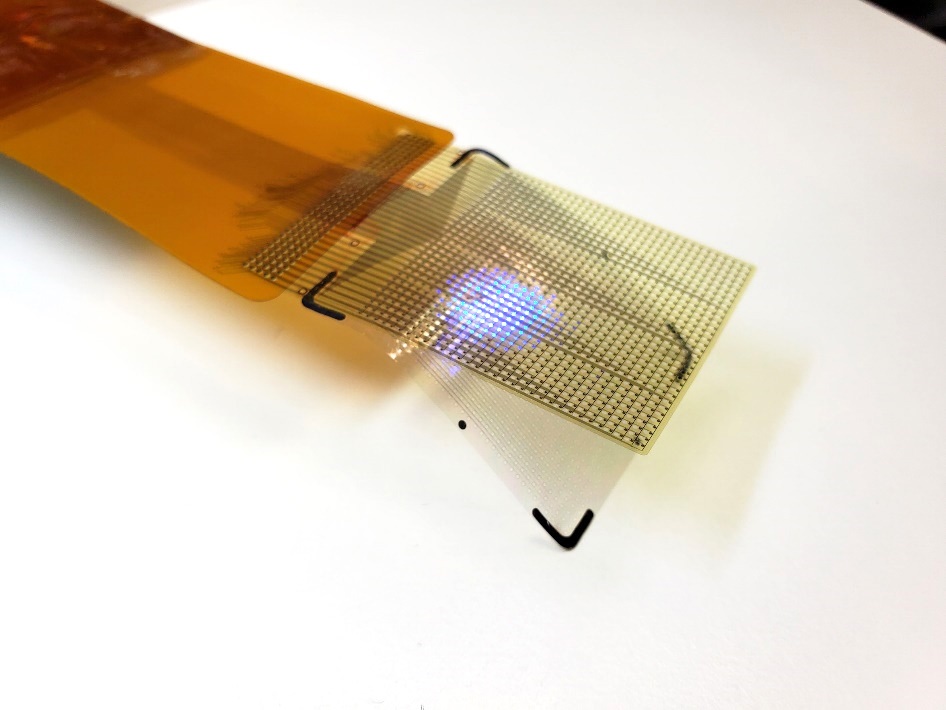
Flexible Microdisplay Visualizes Brain Activity in Real-Time To Guide Neurosurgeons
During brain surgery, neurosurgeons need to identify and preserve regions responsible for critical functions while removing harmful tissue. Traditionally, neurosurgeons rely on a team of electrophysiologists,... Read more.jpg)
Next-Gen Computer Assisted Vacuum Thrombectomy Technology Rapidly Removes Blood Clots
Pulmonary embolism (PE) occurs when a blood clot blocks one of the arteries in the lungs. Often, these clots originate from the leg or another part of the body, a condition known as deep vein thrombosis,... Read morePatient Care
view channel
Surgical Capacity Optimization Solution Helps Hospitals Boost OR Utilization
An innovative solution has the capability to transform surgical capacity utilization by targeting the root cause of surgical block time inefficiencies. Fujitsu Limited’s (Tokyo, Japan) Surgical Capacity... Read more
Game-Changing Innovation in Surgical Instrument Sterilization Significantly Improves OR Throughput
A groundbreaking innovation enables hospitals to significantly improve instrument processing time and throughput in operating rooms (ORs) and sterile processing departments. Turbett Surgical, Inc.... Read more
Next Gen ICU Bed to Help Address Complex Critical Care Needs
As the critical care environment becomes increasingly demanding and complex due to evolving hospital needs, there is a pressing requirement for innovations that can facilitate patient recovery.... Read moreGroundbreaking AI-Powered UV-C Disinfection Technology Redefines Infection Control Landscape
Healthcare-associated infection (HCAI) is a widespread complication in healthcare management, posing a significant health risk due to its potential to increase patient morbidity and mortality, prolong... Read moreHealth IT
view channel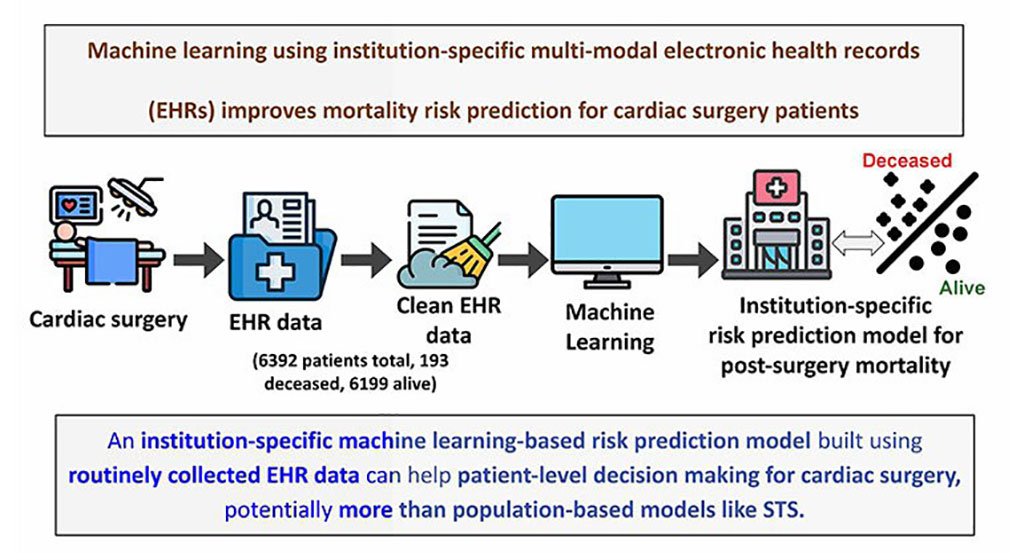
Machine Learning Model Improves Mortality Risk Prediction for Cardiac Surgery Patients
Machine learning algorithms have been deployed to create predictive models in various medical fields, with some demonstrating improved outcomes compared to their standard-of-care counterparts.... Read more
Strategic Collaboration to Develop and Integrate Generative AI into Healthcare
Top industry experts have underscored the immediate requirement for healthcare systems and hospitals to respond to severe cost and margin pressures. Close to half of U.S. hospitals ended 2022 in the red... Read more
AI-Enabled Operating Rooms Solution Helps Hospitals Maximize Utilization and Unlock Capacity
For healthcare organizations, optimizing operating room (OR) utilization during prime time hours is a complex challenge. Surgeons and clinics face difficulties in finding available slots for booking cases,... Read more
AI Predicts Pancreatic Cancer Three Years before Diagnosis from Patients’ Medical Records
Screening for common cancers like breast, cervix, and prostate cancer relies on relatively simple and highly effective techniques, such as mammograms, Pap smears, and blood tests. These methods have revolutionized... Read morePoint of Care
view channel
Critical Bleeding Management System to Help Hospitals Further Standardize Viscoelastic Testing
Surgical procedures are often accompanied by significant blood loss and the subsequent high likelihood of the need for allogeneic blood transfusions. These transfusions, while critical, are linked to various... Read more
Point of Care HIV Test Enables Early Infection Diagnosis for Infants
Early diagnosis and initiation of treatment are crucial for the survival of infants infected with HIV (human immunodeficiency virus). Without treatment, approximately 50% of infants who acquire HIV during... Read more
Whole Blood Rapid Test Aids Assessment of Concussion at Patient's Bedside
In the United States annually, approximately five million individuals seek emergency department care for traumatic brain injuries (TBIs), yet over half of those suspecting a concussion may never get it checked.... Read more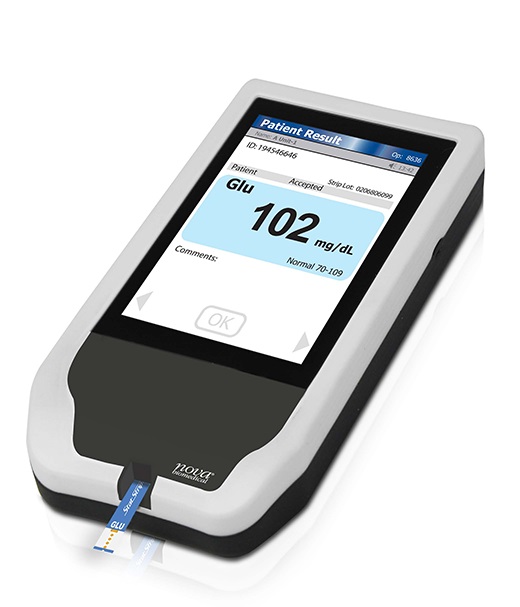
New Generation Glucose Hospital Meter System Ensures Accurate, Interference-Free and Safe Use
A new generation glucose hospital meter system now comes with several features that make hospital glucose testing easier and more secure while continuing to offer accuracy, freedom from interference, and... Read moreBusiness
view channel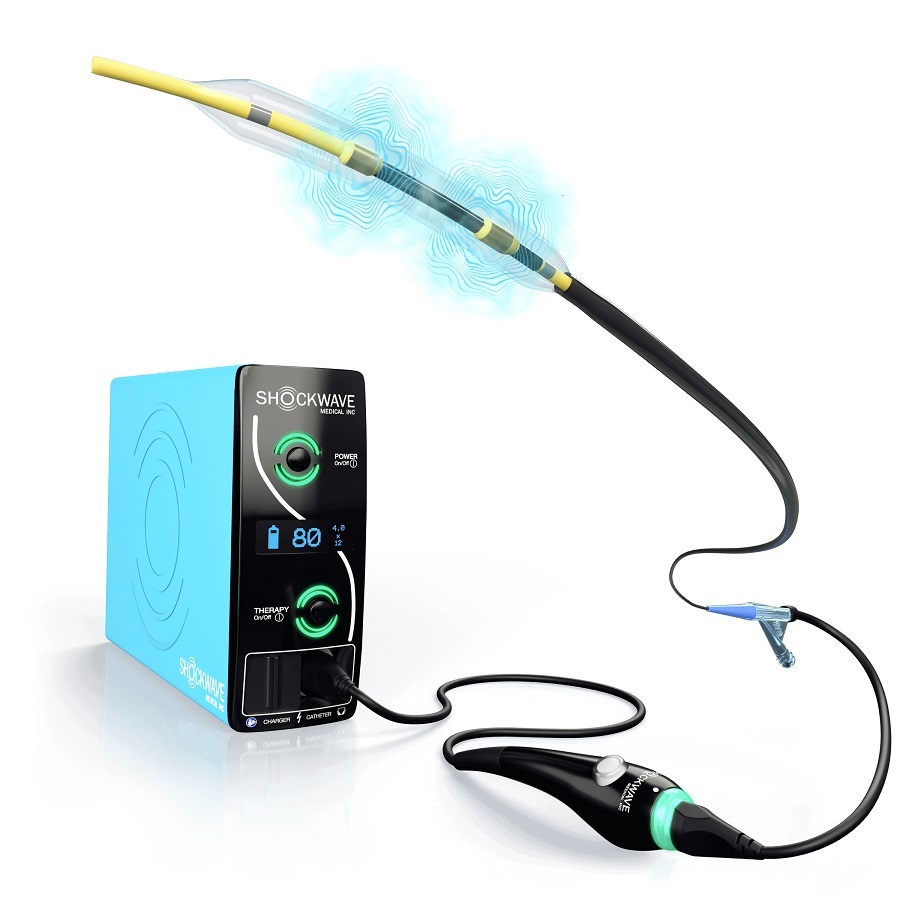
Johnson & Johnson Acquires Cardiovascular Medical Device Company Shockwave Medical
Johnson & Johnson (New Brunswick, N.J., USA) and Shockwave Medical (Santa Clara, CA, USA) have entered into a definitive agreement under which Johnson & Johnson will acquire all of Shockwave’s... Read more